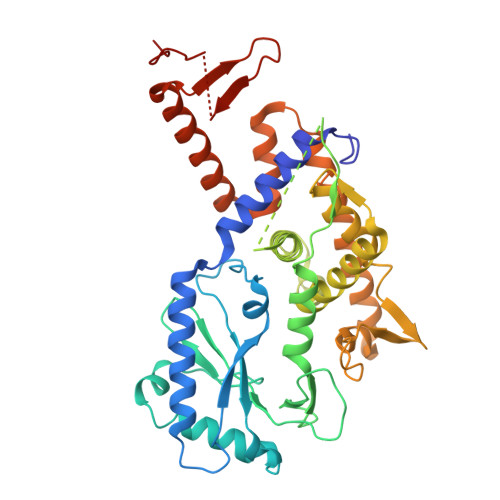
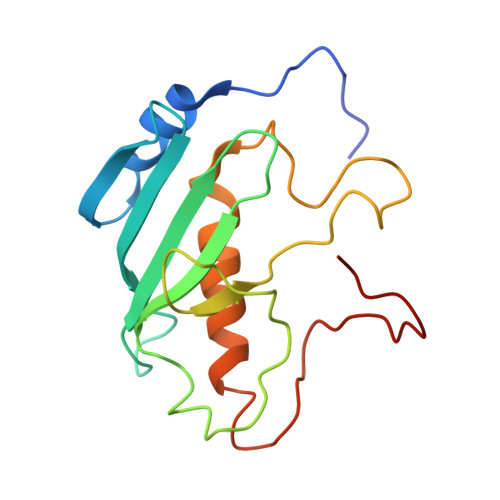
Structures and mechanism of E2-CBASS anti-phage system.
Xiao, J., Yan, Y., Li, J., Oyejobi, G.K., Lan, D., Zhu, B., Wang, Z., Wang, L.(2026) mLife 5: 99-107
- PubMed: 41767953 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/mlf2.70052
- Primary Citation Related Structures:
9KKB - PubMed Abstract:
Bacteria deploy diverse innate immune systems to combat bacteriophage infections. The cyclic-oligonucleotide-based anti-phage signaling system (CBASS) is a type of innate prokaryotic immune system. CBASS synthesizes cyclic-oligonucleotide through cGAS/DncV-like nucleotidyltransferases (CD-NTases) to activate downstream effectors, which kill bacteriophage-infected bacteria, thereby stopping phage spread. One major class of CBASS contains a homolog of eukaryotic ubiquitin-conjugating enzymes, either as an E1-E2 fusion or a single E2 enzyme. Both enzymes function by regulating CD-NTase activity. Currently, many structures of CD-NTases have been reported, but there are only a few reports of structures where CD-NTases form complexes with the associated E2. In this study, we analyzed the length and classification of the CD-NTase in two types of type II CBASS-E1E2/JAB-CBASS and E2-CBASS. We found that the CD-NTase in E2-CBASS is longer and predominantly belongs to clade G. We also present the structure of the Sm CdnG- Sm E2 complex with the bound GTP substrate, which indicates the conservation of the donor binding pattern. Interestingly, we discovered that Sm CdnG contains a conserved C-terminal α-helix and β-sheet structure, which is uniquely involved in forming a complex with Sm E2. We also found that the structure of the E2 protein in the E2-CBASS system is highly conserved. Altogether, we provide mechanistic insights into the E2-CBASS system.
- School of Pharmaceutical Sciences Wuhan University Wuhan China.
Organizational Affiliation: