Structural basis of a sodium pump quaternary complex leading phospholipid scrambling in the living cell
Dou, Y., Abe, K., Suzuki, J.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Sodium/potassium-transporting ATPase subunit alpha-1 | 985 | Homo sapiens | Mutation(s): 0 Gene Names: ATP1A1 EC: 7.2.2.13 | 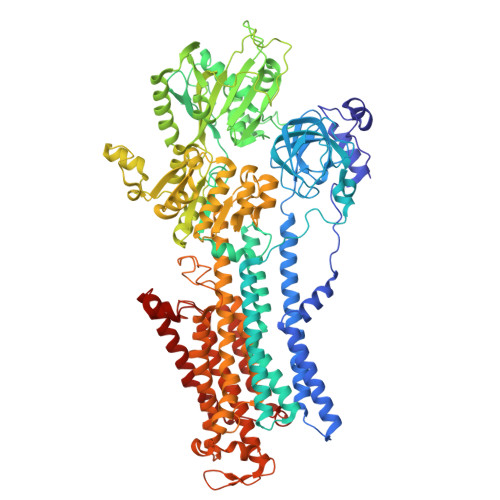 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P05023 GTEx: ENSG00000163399 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P05023 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Sodium/potassium-transporting ATPase subunit beta-3 | 279 | Homo sapiens | Mutation(s): 0 Gene Names: ATP1B3 | 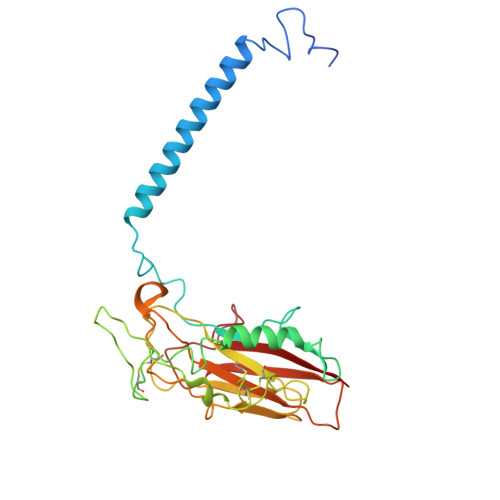 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P54709 GTEx: ENSG00000069849 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P54709 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| FXYD domain-containing ion transport regulator 5 | 178 | Homo sapiens | Mutation(s): 0 Gene Names: FXYD5, DYSAD, IWU1, HSPC113, UNQ2561/PRO6241 | 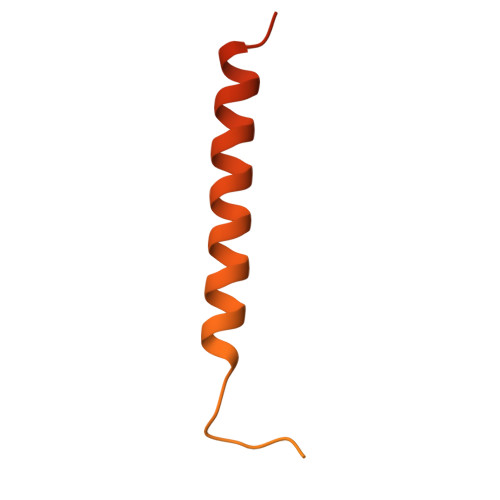 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q96DB9 GTEx: ENSG00000089327 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q96DB9 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 6 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PC1 Download:Ideal Coordinates CCD File | J [auth A], N [auth B] | 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE C44 H88 N O8 P NRJAVPSFFCBXDT-HUESYALOSA-N |  | ||
| PCW Download:Ideal Coordinates CCD File | L [auth A], M [auth B] | 1,2-DIOLEOYL-SN-GLYCERO-3-PHOSPHOCHOLINE C44 H85 N O8 P SNKAWJBJQDLSFF-NVKMUCNASA-O |  | ||
| ACP (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | I [auth A] | PHOSPHOMETHYLPHOSPHONIC ACID ADENYLATE ESTER C11 H18 N5 O12 P3 UFZTZBNSLXELAL-IOSLPCCCSA-N |  | ||
| CLR Download:Ideal Coordinates CCD File | H [auth A], K [auth A], O [auth B] | CHOLESTEROL C27 H46 O HVYWMOMLDIMFJA-DPAQBDIFSA-N |  | ||
| MG Download:Ideal Coordinates CCD File | G [auth A] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| NA (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | D [auth A], E [auth A], F [auth A] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Funding Organization | Location | Grant Number |
|---|---|---|
| Japan Society for the Promotion of Science (JSPS) | Japan | 21H02426 |