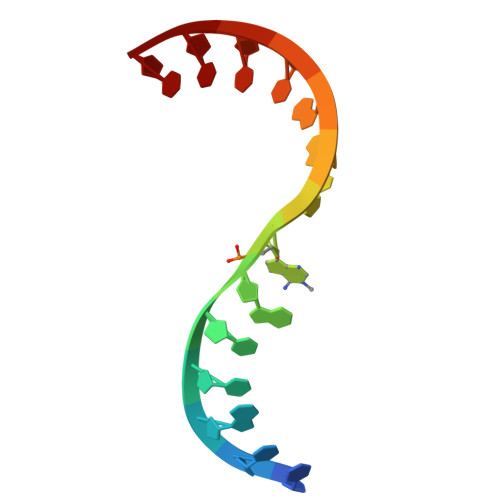
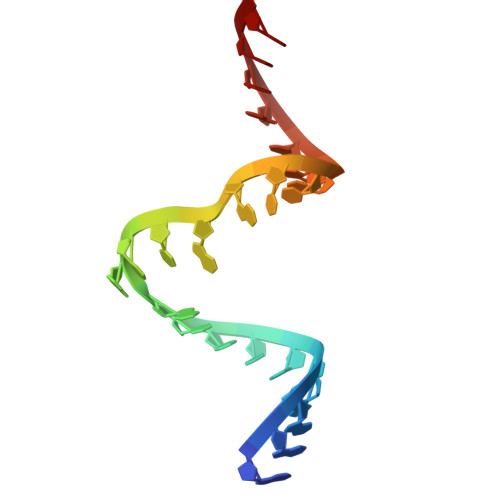
Site-specific ribozyme-mediated alkylation of DNA substrates.
He, Y., Wilson, T.J., Deng, J., Luo, Y., Yan, K., Xie, Y., Lilley, D.M.J., Huang, L.(2026) Nucleic Acids Res 54
- PubMed: 41854073 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkag211
- Primary Citation Related Structures:
9KBR, 9KFV - PubMed Abstract:
MTR1 is an in vitro-selected ribozyme that catalyses the transfer of an alkyl group from exogenous O6-alkylguanine to N1 of a specific adenine in RNA. We show here that the ribozyme can also efficiently alkylate a DNA substrate strand, with almost complete alkylation in 20 min. We have determined crystal structures of the products of methyl and benzyl transfer. The structures are closely similar to that of the all-RNA ribozyme, binding the guanine product in an identical manner, and alkylation occurs at the equivalent location as in the RNA, i.e. N1 of dA63. 2'-O-methylation of C10 and U45, which are hydrogen-bonded to the exogenous guanine, leads to an order-of-magnitude faster rate of alkyl transfer. The results indicate that MTR1 could be a useful tool for the site-specific modification of DNA, including the creation of fluorescent labels or targets for chemical crosslinking.
- Guangdong Provincial Key Laboratory of Malignant Tumor Epigenetics and Gene Regulation, Guangdong-Hong Kong Joint Laboratory for RNA Medicine, Medical Research Center, Sun Yat-Sen Memorial Hospital, Sun Yat-Sen University, Guangzhou 510120, China.
Organizational Affiliation: