A spontaneous termination mechanism of RNA polymerase V shapes the DNA methylation landscape in plants.
Xie, G., Du, X., Tan, Y., Zhou, Y., Chi, C., Zhou, S., Picard, C.L., Chai, S., Wu, L., Zhu, D., Zhao, J., Xue, Y., Li, S., Jacobsen, S.E., Wu, Z., Du, J.(2026) EMBO J 45: 3192-3205
- PubMed: 41928006 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s44318-026-00763-7
- Primary Citation Related Structures:
9K11, 9K12, 9K13, 9K14, 9K15, 9K16, 9K17, 9K18, 9K19 - PubMed Abstract:
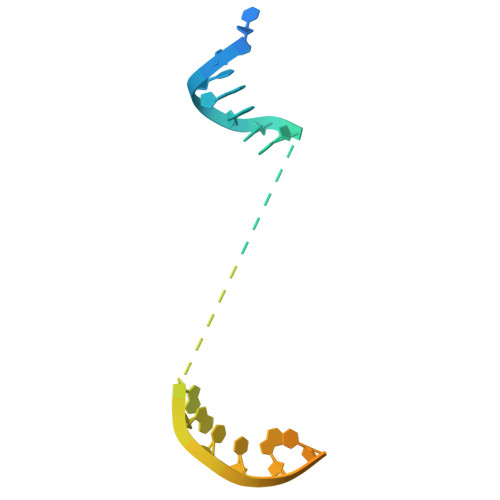
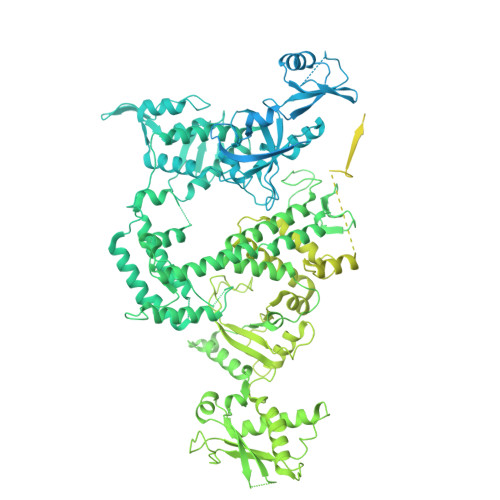
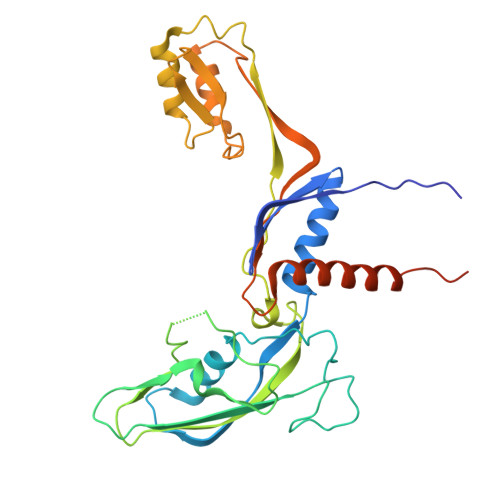
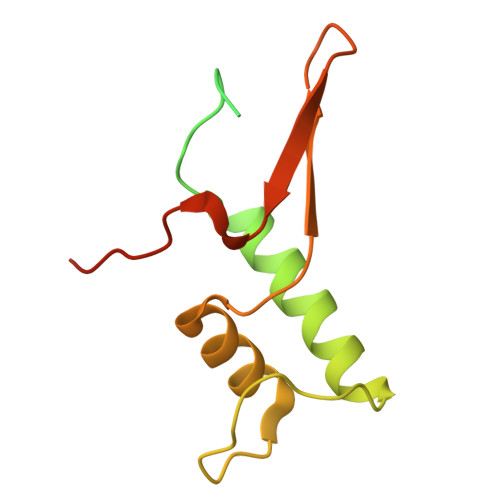
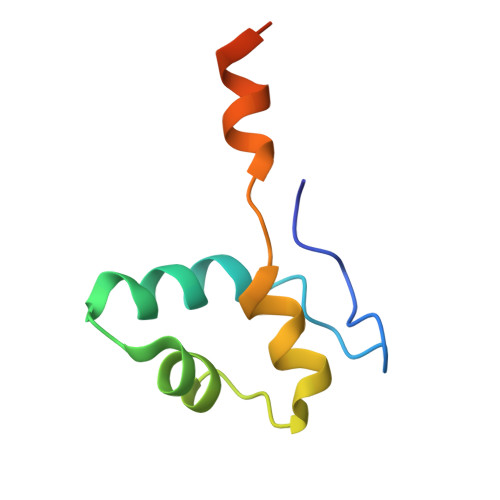
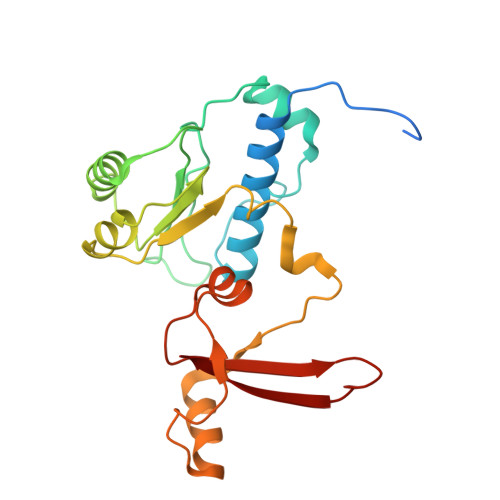
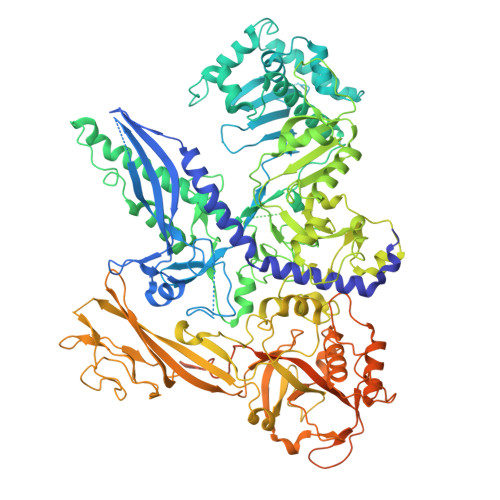
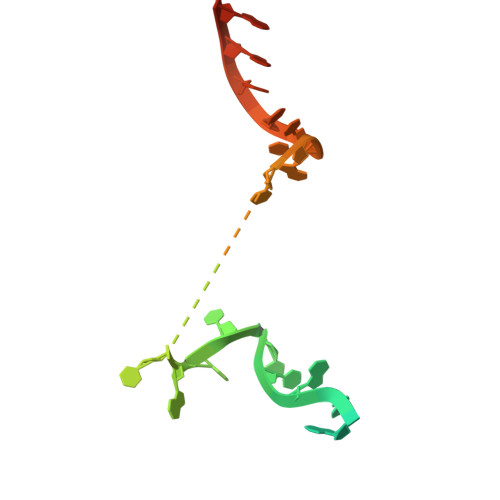
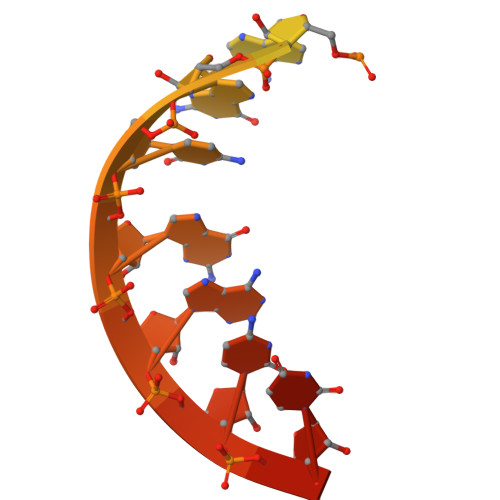
DNA methylation plays critical roles in eukaryotic gene silencing, genome imprinting, viral defense, and suppression of transposable elements. In plants, RNA Polymerase V (Pol V)-generated non-coding RNA guides DNA methylation through the RNA-directed DNA methylation (RdDM) pathway; however, how these RNAs are selected is unknown. Here, we show that the 3'-ends of Pol V transcripts are enriched at A-rich template DNA (A-rich-DNA T ). Arabidopsis RdDM regions possess AT-rich boundaries genome-wide, suggesting that Pol V likely terminates at A-rich-DNA T , which subsequently defines the DNA methylation landscape in plants. A-rich-DNA T successfully stops Pol V transcription in vitro. Structural snapshots of Pol V transcribing A-rich-DNA T show that accumulation of unstable rU:dA pairs in the RNA-DNA hybrid promotes transcription bubble collapse and spontaneous transcription termination. These findings identify an intrinsic Pol V termination signal that shapes genomic DNA methylation patterning in plants and reveals a common mechanism for spontaneous transcription termination.
- Shenzhen Key Laboratory of Plant Genetic Engineering and Molecular Design, Institute of Plant and Food Science, Department of Biology, School of Life Sciences, Southern University of Science and Technology, Shenzhen, China.
Organizational Affiliation: