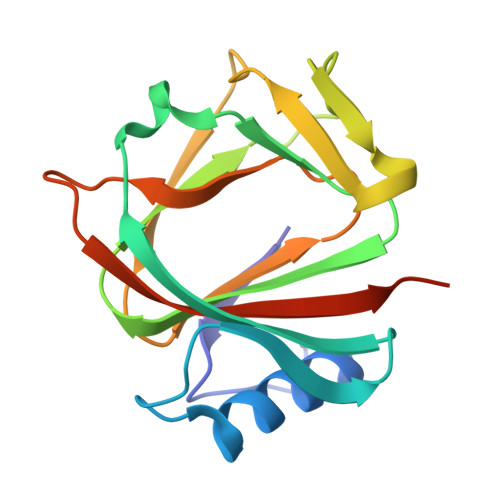
Crystal Structure and Functional Characterization of YjgK From Salmonella Typhimurium.
Choi, S.Y., Kim, E., Yoon, H., Kim, H.N., Kim, J.H., Seok, S.H., Seo, M.D.(2025) Proteins 93: 1935-1943
- PubMed: 40525813 Search on PubMed
- DOI: https://doi.org/10.1002/prot.26854
- Primary Citation Related Structures:
9JRR - PubMed Abstract:
The YhcH/YjgK/YiaL (DUF386) family, widely conserved across bacterial species, is involved in essential cellular processes yet remains poorly characterized. YjgK from Salmonella enterica serovar Typhimurium has drawn attention because of its potential role in biofilm formation associated with metal homeostasis, which may be critical for bacterial survival. In this study, we report the crystal structure of YjgK at 1.76 Å resolution, revealing a dimeric arrangement where each monomer consists of a jelly roll-type β-sandwich fold. This fold forms a funnel-shaped cavity, suggesting potential ligand binding. YjgK contains two zinc ions per dimer, which were identified through structural analysis and confirmed by inductively coupled plasma mass spectrometry (ICP-MS). The zinc ions are coordinated by conserved residues (Glu62, His64, Asp69, and His128) to form a tetrahedral geometry. Structural comparisons with homologous proteins revealed significant similarities in their overall fold but distinct differences in their metal ion specificity, with YhcH binding copper and HP1029 binding zinc. Salmonella lacking YjgK increased biofilm formation, while YjgK overexpression hardly influenced biofilm formation. Our findings suggest that the zinc-binding capability of YjgK may play a key role in metal ion homeostasis, contributing to the ability of Salmonella to form biofilm in response to metal-limited environments, such as those encountered during infection. The conservation of DUF386 fold across species, along with variations in metal ion coordination, indicates functional diversification within this family.
- College of Pharmacy and Research Institute of Pharmaceutical Science and Technology (RIPST), Ajou University, Suwon, Republic of Korea.
Organizational Affiliation: