Classical conformation for the spCas9 with gRNA and target DNA complex
Xi, Z.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| CRISPR-associated endonuclease Cas9/Csn1 | C [auth A] | 1,368 | Streptococcus pyogenes serotype M1 | Mutation(s): 0 Gene Names: cas9 EC: 3.1 | 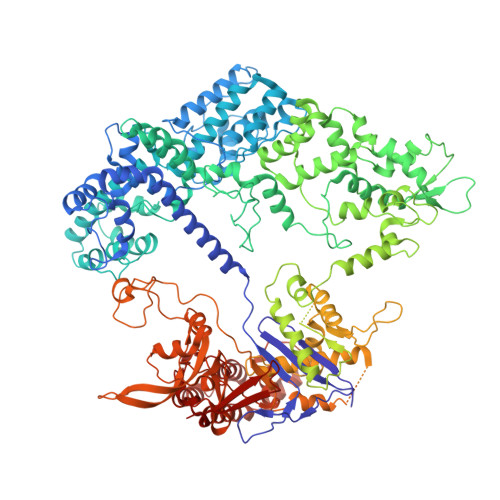 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q99ZW2 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 1 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| TS | A [auth C] | 28 | unclassified sequences |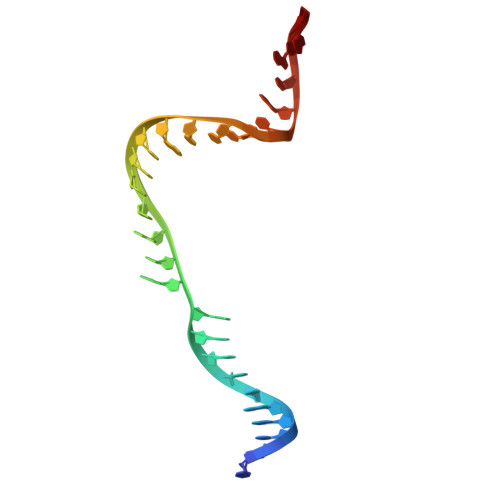  |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
Entity ID: 2 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| NTS | B [auth D] | 28 | unclassified sequences |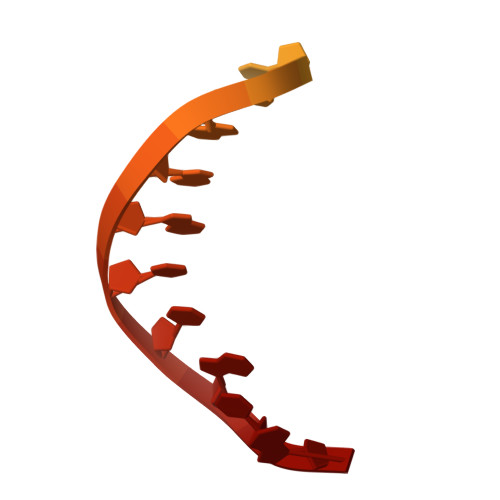  |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
Entity ID: 4 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| gRNA | D [auth B] | 98 | unclassified sequences | 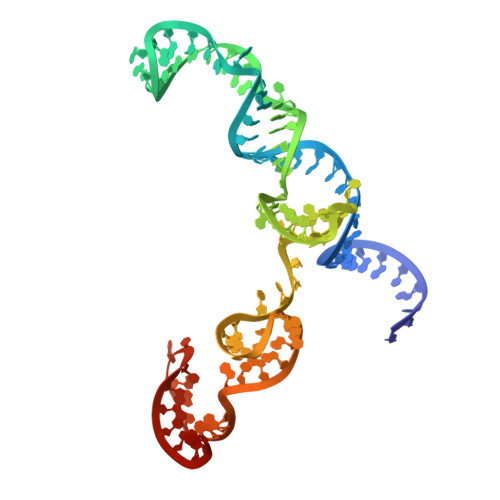 |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Science Foundation (NSF, China) | China | -- |