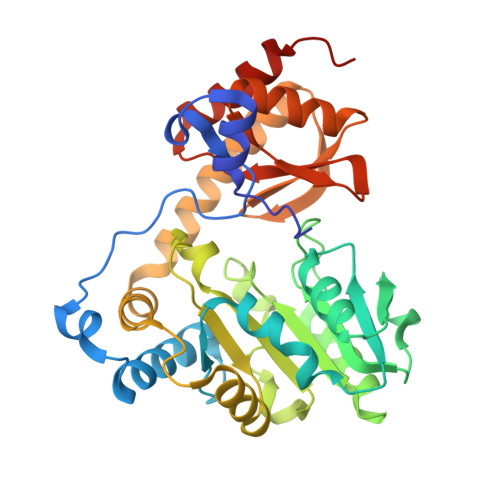
Crystal structure of l-threonine-O-3-phosphate decarboxylase CobC from Sinorhizobium meliloti involved in vitamin B 12 biosynthesis.
Jiang, M., Guo, S., Chen, X., Wei, Q., Wang, M.(2024) Biochem Biophys Res Commun 734: 150767-150767
- PubMed: 39366178 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2024.150767
- Primary Citation Related Structures:
9JFB, 9JFF - PubMed Abstract:
Vitamin B 12 is involved in many important biochemical reactions for humans, and its deficiency can lead to serious diseases. The industrial production of vitamin B 12 is achieved through microbial fermentation. In this work, we determine the crystal structures of the l-threonine-O-3-phosphate (Thr-P) decarboxylase CobC from Sinorhizobium meliloti (SmCobC), an industrial vitamin B 12 -producing bacterium, in apo form and in complex with a reaction intermediate. Our structures supported the Thr-P decarboxylase activity of SmCobC and revealed that the positively charged substrate-binding pocket between the large and small domains determines its substrate selectivity for Thr-P. Moreover, our results provided evidence for the proposition that the AP-P linker is formed by direct incorporation of AP-P in the biosynthetic pathway of vitamin B 12 in S.meliloti.
- Institutes of Physical Science and Information Technology, Anhui University, Hefei, 230601, Anhui, China; School of Life Sciences, Anhui University, Hefei, 230601, Anhui, China.
Organizational Affiliation: