Structural and functional characterization of CspR, a 2'-O-methyltransferase acting on wobble position within tRNA.
Yoo, J., Lee, Y., Cho, G., Lim, J., Kim, J.(2025) Nucleic Acids Res 53
- PubMed: 40794868 Search on PubMed
- DOI: https://doi.org/10.1093/nar/gkaf751
- Primary Citation Related Structures:
8W9U, 9J5P - PubMed Abstract:
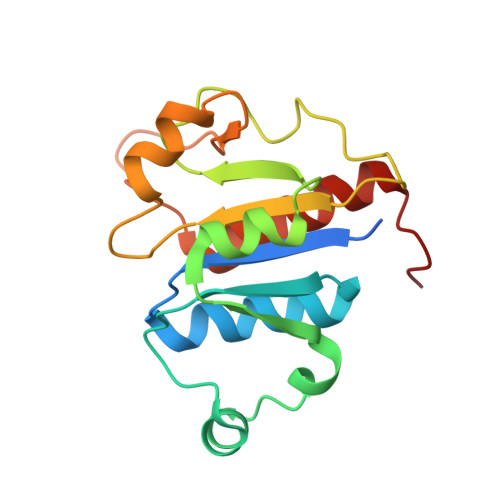
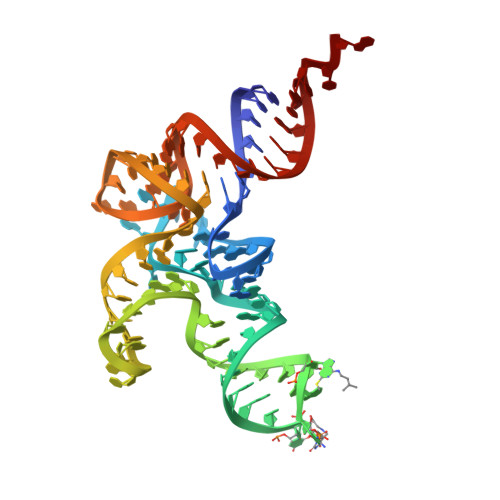
Post-transcriptional modifications of transfer RNA (tRNA) are essential for maintaining decoding fidelity and tRNA stability. Among these, 2'-O-ribosyl methylation is particularly prominent in bacteria. In this study, we provide structural and biochemical evidence identifying CspR from Bacillus subtilis as an ortholog of Escherichia coli TrmL, which specifically methylates the 2'-O-ribose of the wobble position of tRNALeu(CAA), tRNALeu(UAA), and tRNAPhe(GAA), in the presence of 2-methylthio-N6-isopentenyladenosine at position 37 (ms2i6A37). X-ray crystal structures of CspR in complex with tRNALeu(UAA) and in its tRNA-free form reveal substantial conformational rearrangements upon tRNA binding. Notably, the tRNA-bound structure shows specific interactions between the anticodon loop and CspR's dimeric interface, with key residues U33, 5-carboxymethylaminomethyluridine34 (cmnm5U34), and ms2i6A37 adopting flipped-out conformations. Furthermore, the structure uncovers extensive hydrophobic interactions between the isopentenyl group of ms2i6A37 and CspR, explaining the critical requirement of hypermodified A37 for enzymatic activity.
- Department of Chemistry, Gwangju Institute of Science and Technology, Gwangju 61005, Korea.
Organizational Affiliation: