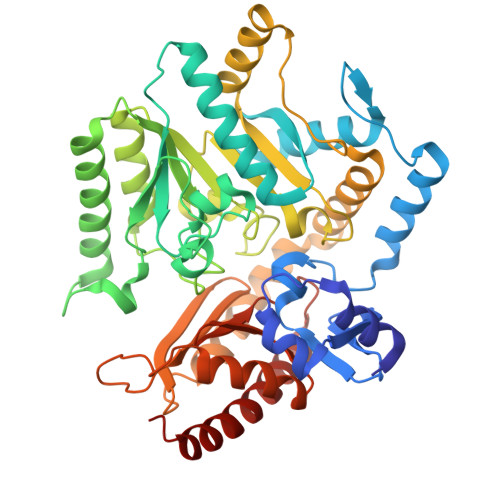
Structural insights and rational design of Pseudomonasputida KT2440 omega transaminases for enhanced biotransformation of (R)-PAC to (1R, 2S)-Norephedrine.
Das, P., Noronha, S., Bhaumik, P.(2025) J Biological Chem 301: 110289-110289
- PubMed: 40436318 Search on PubMed
- DOI: https://doi.org/10.1016/j.jbc.2025.110289
- Primary Citation Related Structures:
9J2K, 9J4Y, 9J4Z, 9J50 - PubMed Abstract:
Omega transaminases (ω-TAs) can mediate the chiral amination of several unnatural substrates without the requirement of an α-COOH group and are highly relevant in the production of several pharmaceutical intermediates of commercial interest. Development of better variants of ω-TAs is hence essential for biotransformation of unnatural substrates. We studied the active site architecture of the wild-type ω-TAs, to engineer enzymes that enhance the biotransformation of (R)-phenylacetylcarbinol to (1R, 2S)-norephedrine. Two such ω-TAs (TA_5182 and TA_2799) from P. putida KT2440 strain were overexpressed and purified as recombinant proteins. Crystal structures of TA_5182 were solved in two conformations, and significant movements of two highly flexible loops were observed in these different states. The TA_2799 structure was determined as a complex with the cofactor pyridoxal 5'-phosphate (PLP) covalently bound to the catalytic K286 as an internal aldimine. Enzyme assays indicated that TA_2799 required a four-fold higher cofactor concentration than TA_5182 to achieve satisfactory biotransformation of (R)-PAC. A key mutation of L322F in TA_2799 drastically reduced (∼8-fold) the cofactor dependency of the TA_2799_L322F mutant enzyme, and the mutant remained active for 96 h at 30°C. The crystal structure of the mutant enzyme revealed a key asparagine residue that mediates a hydrogen bonding network at the dimeric interface of the enzyme and is absent in TA_5182. The TA_5182_G119N mutant also showed enhanced cofactor affinity. The results of our studies will help generate Pseudomonad ω-TAs and ω-TAs from other organisms with high efficiency for asymmetric synthesis, for further applications in large-scale biotransformation processes.
- Department of Biosciences and Bioengineering, Indian Institute of Technology Bombay, Powai, Mumbai-400076, India.
Organizational Affiliation: