VLP structure of Chikungunya virus complexed with C37 Fab, 2f block.
Han, X., Ji, C., Wang, F., Tian, S., Gao, F.G., Yan, J.To be published.
Experimental Data Snapshot
Starting Models: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| C37 Fab, Heavy chain | A [auth S], O [auth Q], Q [auth O], S [auth M] | 265 | Homo sapiens | Mutation(s): 0 | 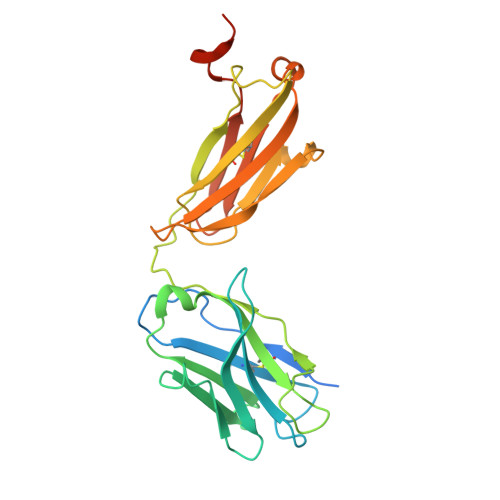 |
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| C37 Fab, Light chain | B [auth T], P [auth R], R [auth P], T [auth N] | 241 | Homo sapiens | Mutation(s): 0 | 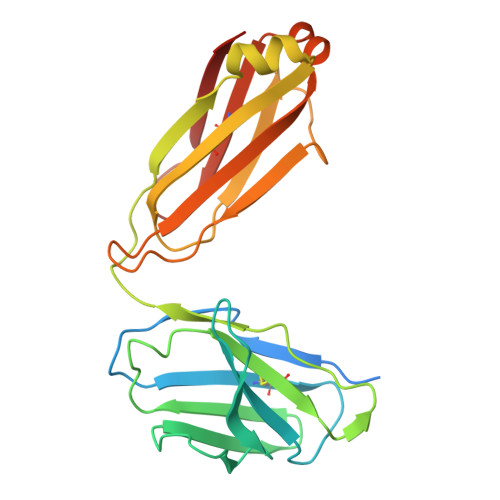 |
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| CHIKV E1 | C [auth A], D [auth B], E [auth C], F [auth D] | 439 | Chikungunya virus | Mutation(s): 0 | 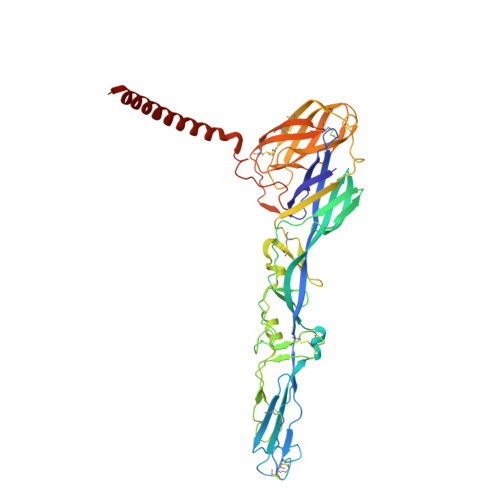 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A286S4J4 | ||||
Glycosylation | |||||
| Glycosylation Sites: 1 | |||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Structural polyprotein | G [auth E], H [auth F], I [auth G], J [auth H] | 419 | Chikungunya virus | Mutation(s): 0 Gene Names: CHIKVgp2 | 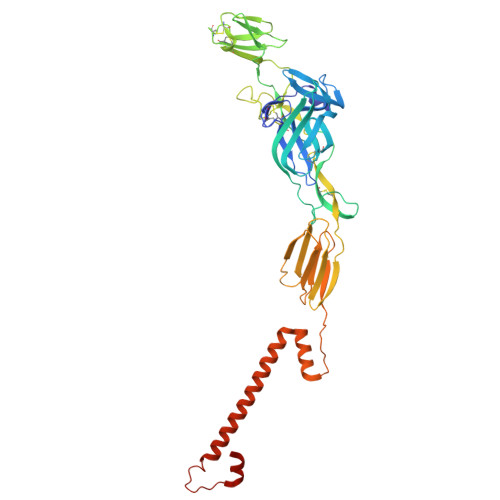 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | D7R9A2 | ||||
Glycosylation | |||||
| Glycosylation Sites: 2 | |||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| CHIKV capsid protein | K [auth I], L [auth J], M [auth K], N [auth L] | 151 | Chikungunya virus | Mutation(s): 0 EC: 3.4.21.90 |  |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A6B9KBE1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 6 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose | U, V, X | 4 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G32152BH GlyCosmos: G32152BH GlyGen: G32152BH | |||||
Entity ID: 7 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| alpha-D-mannopyranose-(1-3)-beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose | W | 5 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G42466VF GlyCosmos: G42466VF GlyGen: G42466VF | |||||
Entity ID: 8 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | CA [auth c], EA [auth e], Y | 5 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G10133VD GlyCosmos: G10133VD GlyGen: G10133VD | |||||
Entity ID: 9 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | BA [auth b], DA [auth d], Z | 3 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G15407YE GlyCosmos: G15407YE GlyGen: G15407YE | |||||
Entity ID: 10 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| alpha-D-mannopyranose-(1-3)-beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | AA [auth a] | 4 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G81315DD GlyCosmos: G81315DD GlyGen: G81315DD | |||||
Entity ID: 11 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| alpha-D-mannopyranose-(1-6)-beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | FA [auth f] | 4 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G22573RC GlyCosmos: G22573RC GlyGen: G22573RC | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| CLR Download:Ideal Coordinates CCD File | GA [auth B], HA [auth D], IA [auth E], JA [auth G] | CHOLESTEROL C27 H46 O HVYWMOMLDIMFJA-DPAQBDIFSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Natural Science Foundation of China (NSFC) | China | -- |