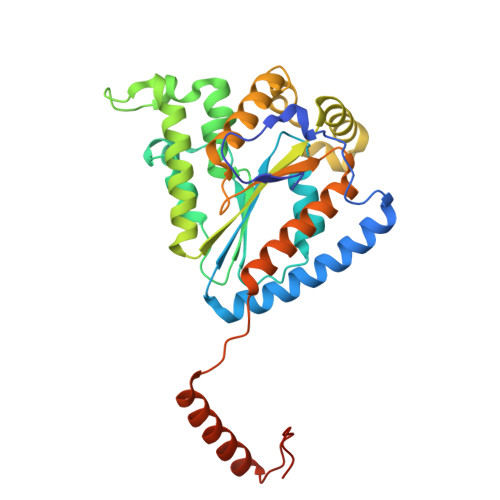
Structure-guided engineering of a polyphosphate kinase 2 class III from an Erysipelotrichaceae bacterium to produce base-modified purine nucleotides.
Mitton-Fry, R.M., Rasche, R., Lawrence-Dorner, A.M., Eschenbach, J., Tekath, A., Rentmeister, A., Kummel, D., Cornelissen, N.V.(2025) RSC Chem Biol 6: 1328-1335
- PubMed: 40667417 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/d5cb00108k
- Primary Citation Related Structures:
9IGQ, 9IGR - PubMed Abstract:
Nucleobase-modified nucleoside-5'-triphosphates (NTPs) are important building blocks for the enzymatic synthesis of non-coding RNAs and mRNAs with improved properties. Chemical phosphorylation of base-modified nucleotides to NTPs remains challenging. Here, we report the enzymatic phosphorylation of purine-modified nucleoside-5'-monophosphates (NMPs) to the corresponding NTPs by the polyphosphate kinase 2 class III from an Erysipelotrichaceae bacterium (EbPPK2). The enzyme is highly promiscuous, accepting a range of NMPs with purine modifications. EbPPK2 efficiently catalyses the formation of the corresponding di-, tri- and tetraphosphates, typically with >70% conversion to the NTP. Slower conversion was observed for analogues with oxo- or thio-substitutions at the C6-position. To better understand nucleotide binding and catalysis, we determined the crystal structure of EbPPK2 at 1.7 Å resolution bound to a non-hydrolysable ATP analogue and polyphosphate. This enabled structure-guided design of EbPPK2 variants that efficiently convert GMP analogues, while retaining activity for AMP. Apart from being the preferred industrial-scale ATP recycling catalyst, EbPPK2 and variants bear potential to become the favoured enzyme family for purine-modified NTP production.
- Institute of Biochemistry, University of Münster, Corrensstr. 36 D-48149 Münster Germany cornelissen@uni-muenster.de.
Organizational Affiliation: