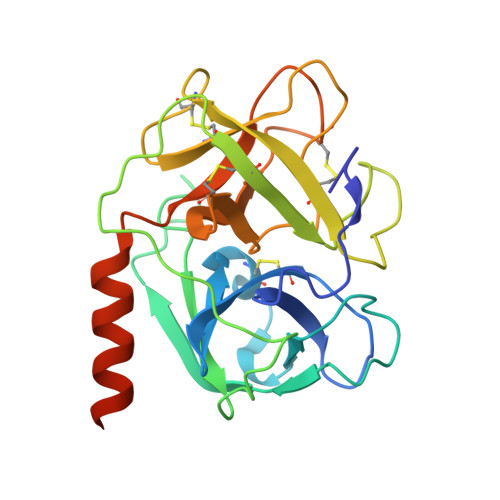
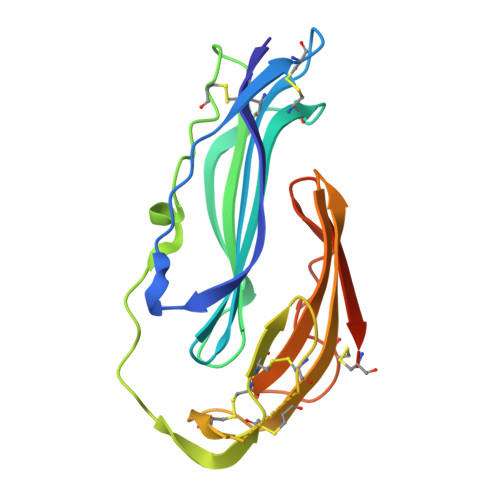
Structures of proteinase 3 and the CD177 receptor complex reveal a major autoantibody epitope.
Zheng-Gerard, C., Joha, J., Carrasquero, M., El Omari, K., Lowe, E., Dubey, S., Draper, S.J., Chang, Y.C., Lin, H.H., Salama, A.D., McHugh, K., Seiradake, E.(2026) EMBO Rep 27: 1580-1606
- PubMed: 41703070 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s44319-026-00716-5
- Primary Citation Related Structures:
9IGO, 9IGP - PubMed Abstract:
Granulomatosis with polyangiitis is a life-threatening systemic vasculitis, characterised by anti-neutrophil cytoplasmic autoantibodies (ANCA) most commonly against proteinase 3 (PR3), a protease expressed intracellularly and on the surface of neutrophils. Most cell surface PR3 is bound to the receptor CD177; however, the molecular mechanism of the interactions is not well understood. Here, we present crystal structures of CD177 in complex with PR3 and unliganded CD177. We describe a mainly hydrophobic binding interface between PR3 and CD177, involving the first two Ly6/uPAR (LU) domains of CD177. These form a globular structure which is connected to downstream domains via a flexible linker. Using a panel of PR3-ANCA-positive patient samples, we show that a significant proportion of ANCAs target the CD177-binding site of PR3 in these samples. Structure-guided mutation of the CD177-binding site on PR3 is effective in reducing PR3-ANCA binding. The results demonstrate that the CD177-binding surface of PR3 harbours a major PR3-ANCA epitope, and that the extent of binding to this surface varies between different patients.
- Department of Biochemistry, University of Oxford, South Parks Road, Oxford, OX1 3QU, UK.
Organizational Affiliation: