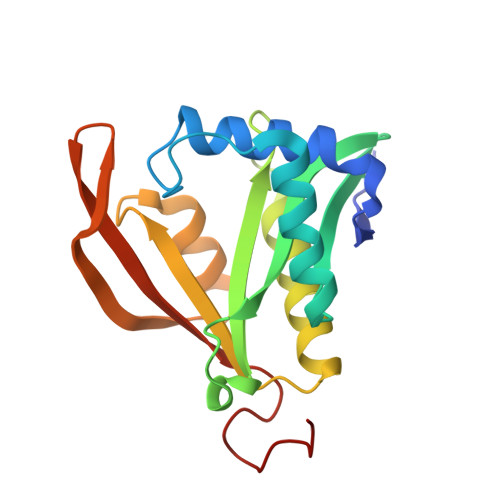
Structure and activity of a phosphinothricin N-acetyltransferase (PSPTO_3321) from Pseudomonas syringae pv. tomato DC3000.
Davies, A.M., Trentham, D., Sutton, B.J., Brown, P.R.(2025) Biochem Biophys Res Commun 755: 151539-151539
- PubMed: 40054337 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2025.151539
- Primary Citation Related Structures:
9IGK, 9IGL - PubMed Abstract:
Phosphinothricin inhibits plant glutamine synthetase and is used as a herbicide. Streptomyces hygroscopicus and Streptomyces viridochromogenes, which produce phosphinothricin naturally, encode acetyltransferases that confer phosphinothricin resistance. In the Pseudomonas genome database, a number of proteins have been annotated as phosphinothricin acetyltransferases and putative phosphinothricin acetyltransferases. One such protein is PSPTO_3321 from P. syringae, a strain that causes tomato speck. Here, we reveal that PSPTO_3321 from P. syringae, termed syr_pat, is a phosphinothricin acetyltransferase, and also retains a lower level of activity against the structurally similar substrate methionine sulfoximine. We solved a 1.6 Å resolution crystal structure of syr_pat alone and a 2.5 Å resolution structure for a complex with L -phosphinothricin. We also characterised active site mutants, providing insights into substrate specificity. Our work now provides a basis for further study of the reaction mechanism.
- King's College London, Randall Centre for Cell and Molecular Biophysics, New Hunt's House, London, SE1 1UL, United Kingdom.
Organizational Affiliation: