Modelling POLG mutations in mice unravels a critical role of POL gamma Beta in regulating phenotypic severity.
Corra, S., Zuppardo, A., Valenzuela, S., Jenninger, L., Cerutti, R., Sillamaa, S., Hoberg, E., Johansson, K.A.S., Rovsnik, U., Volta, S., Silva-Pinheiro, P., Davis, H., Trifunovic, A., Minczuk, M., Gustafsson, C.M., Suomalainen, A., Zeviani, M., Macao, B., Zhu, X., Falkenberg, M., Viscomi, C.(2025) Nat Commun 16: 4782-4782
- PubMed: 40404629 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-60059-y
- Primary Citation Related Structures:
9G74, 9G75, 9G77, 9IBX, 9IBZ, 9IC0, 9IC1, 9IC3 - PubMed Abstract:
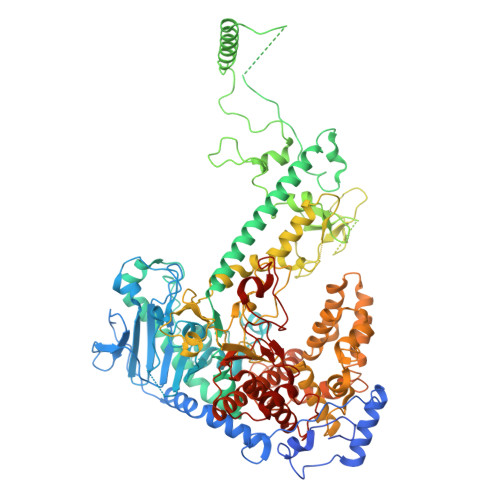
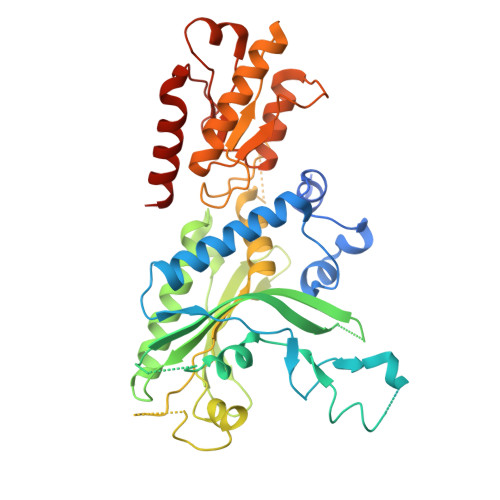
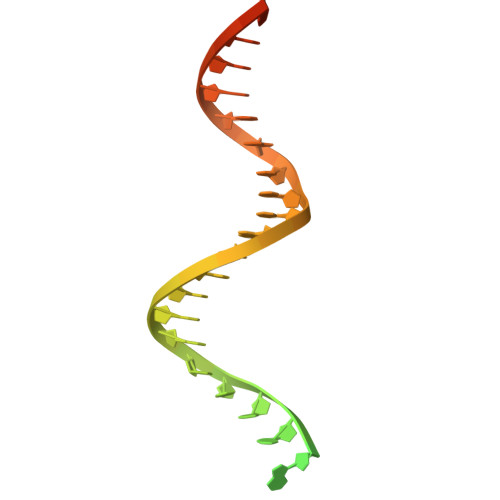
DNA polymerase γ (POLγ), responsible for mitochondrial DNA replication, consists of a catalytic POLγA subunit and two accessory POLγB subunits. Mutations in POLG, which encodes POLγA, lead to various mitochondrial diseases. We investigated the most common POLG mutations (A467T, W748S, G848S, Y955C) by characterizing human and mouse POLγ variants. Our data reveal that these mutations significantly impair POLγ activities, with mouse variants exhibiting milder defects. Cryogenic electron microscopy highlighted structural differences between human and mouse POLγ, particularly in the POLγB subunit, which may explain the higher activity of mouse POLγ and the reduced severity of mutations in mice. We further generated a panel of mouse models mirroring common human POLG mutations, providing crucial insights into the pathogenesis of POLG-related disorders and establishing robust models for therapeutic development. Our findings emphasize the importance of POLγB in modulating the severity of POLG mutations.
- Veneto Institute of Molecular Medicine (VIMM), Via Orus, 2-35129, Padova, Italy.
Organizational Affiliation: