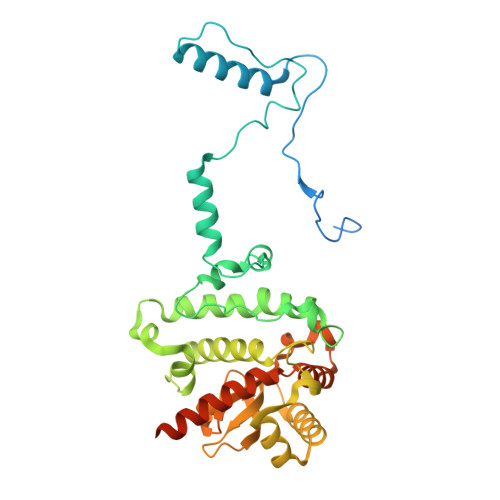
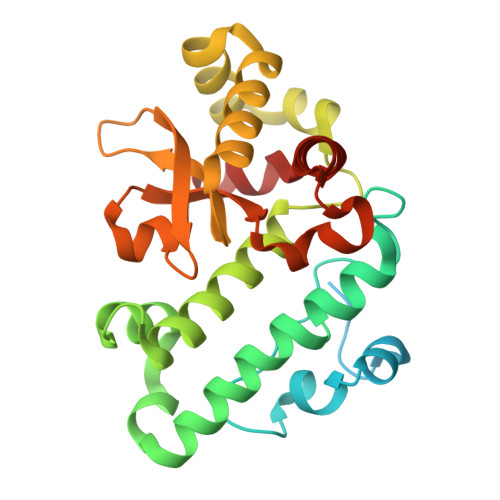
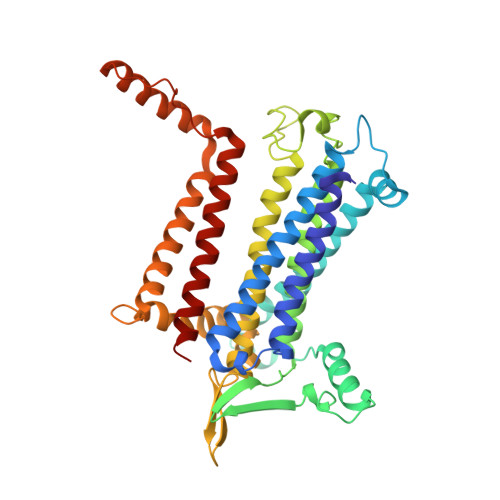
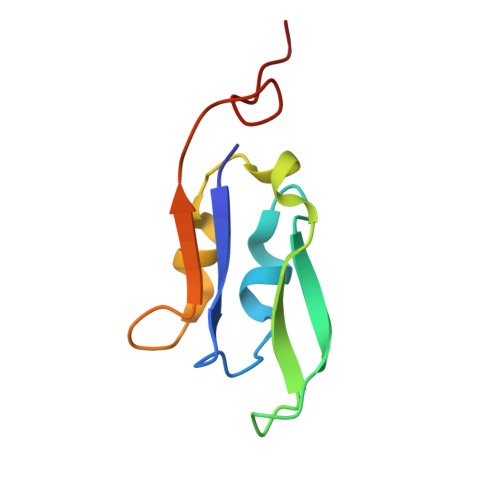
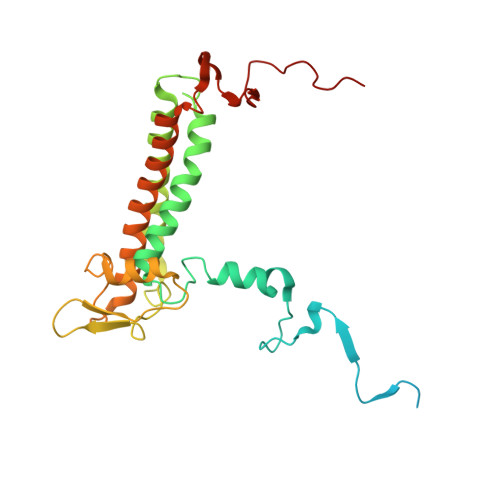
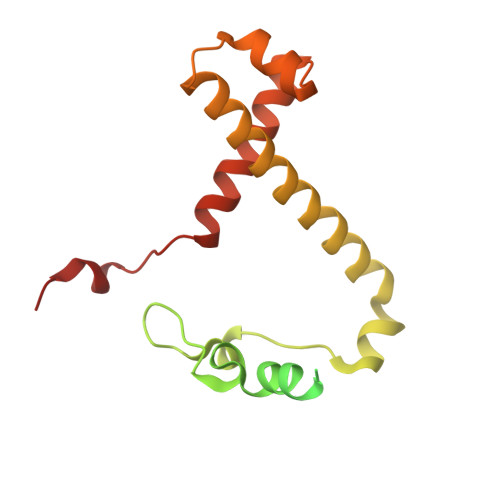
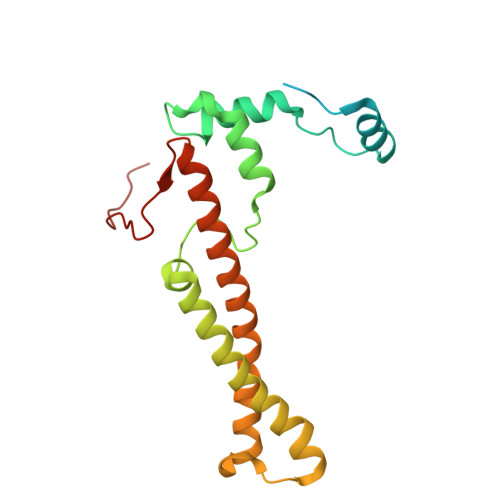
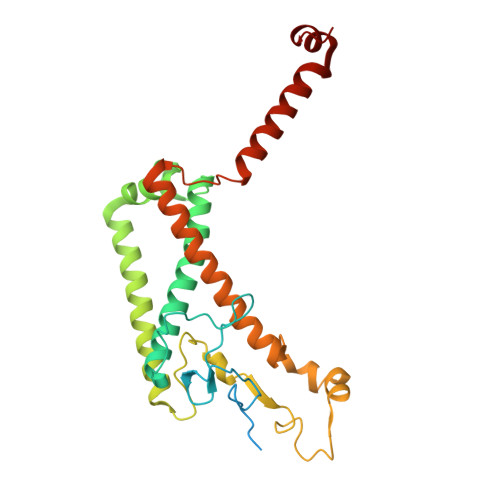
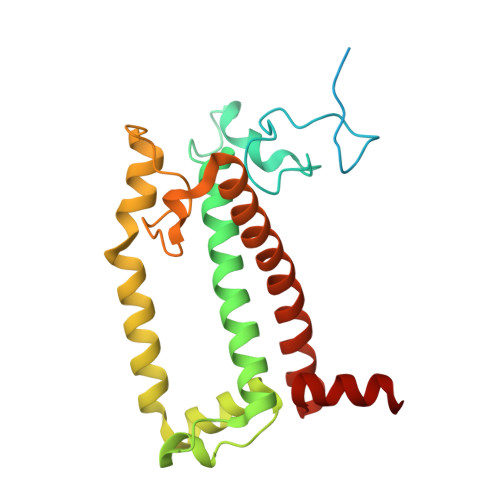
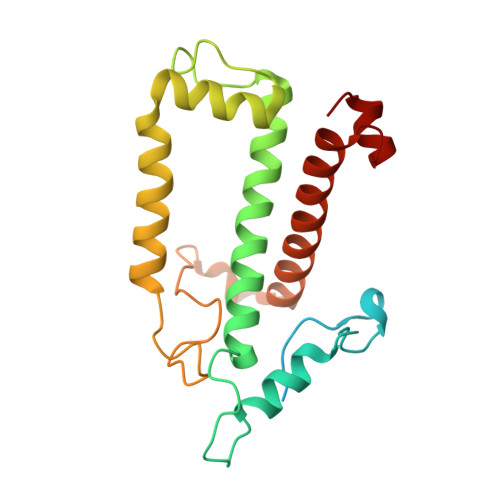
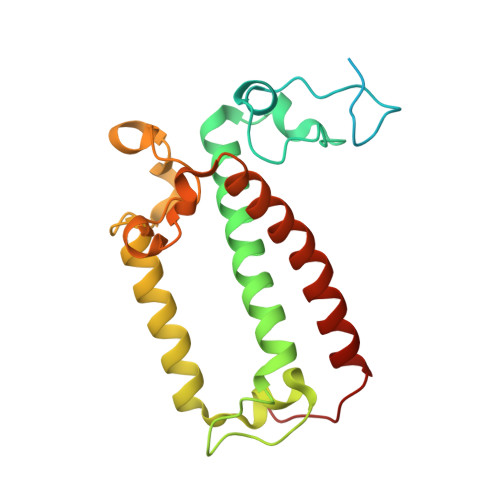
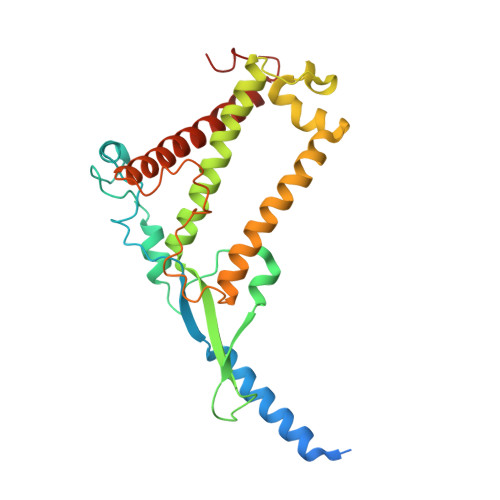
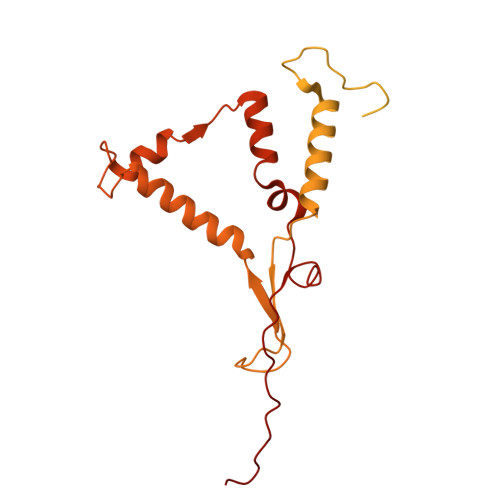
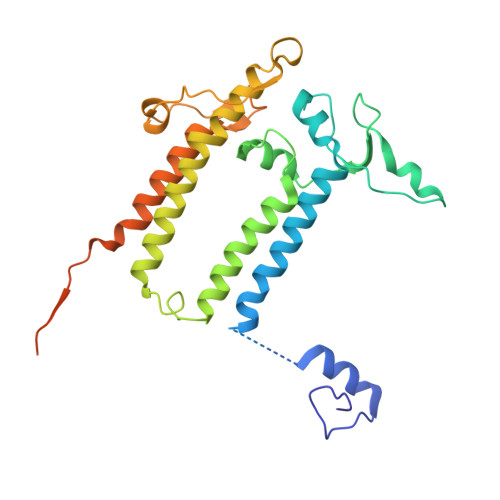
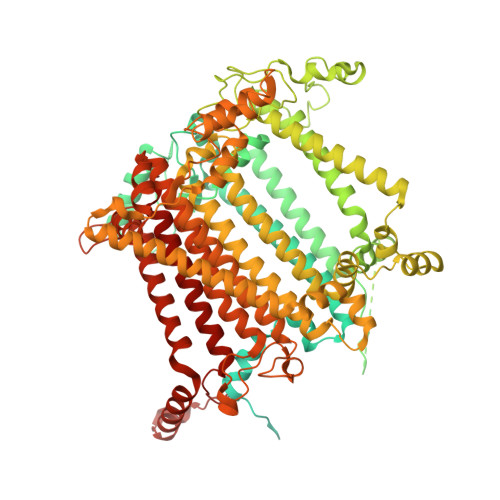
The cryo-EM structure of Photosystem I from Chromera velia with a bound superoxide dismutase heterodimer.
Yuan, X., Lukes, M., Qian, P., Charras, Q., Smrcka, T., Man, P., Kovarova, L., Konik, P., Heilmann, E., Al-Amoudi, A., Sobotka, R., Naschberger, A.(2025) Nat Commun 17: 913-913
- PubMed: 41423677 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-67637-0
- Primary Citation Related Structures:
9HYU - PubMed Abstract:
Photosystem I is a key component of the solar energy conversion machinery in oxygenic photosynthesis, and its core, where photochemistry occurs, is highly conserved. However, the coral-associated alga Chromera velia that is evolutionary linked to parasitic apicomplexans, exhibits Photosystem I with unusual features. These include the splitting of the central PsaA subunit and the binding of superoxide dismutases as regular subunits. The organization of such a unique Photosystem I was enigmatic. Here, we present the cryo-EM structure of Chromera velia Photosystem I at 1.84 Å resolution. Our work reveals a superoxide dismutase heterodimer bound to the stromal side of the core, stabilized by extensions of canonical subunits, a novel protein PsaV, and a reduced light-harvesting apparatus. We elucidate how the complex evolved to accommodate the superoxide dismutase, assemble the split PsaA, and integrate antenna proteins in a non-canonical orientation. Based on our data and prior physiological data, we propose that this specialized Photosystem I functions likely as an Mehler machine, redirecting electrons from Photosystem II back to water. This mechanism enables Chromera velia to manage redox imbalance and reduce photorespiration through localized oxygen consumption.
- King Abdullah University of Science and Technology (KAUST), Biological and Environmental Science and Engineering Division, Thuwal, Saudi Arabia.
Organizational Affiliation: