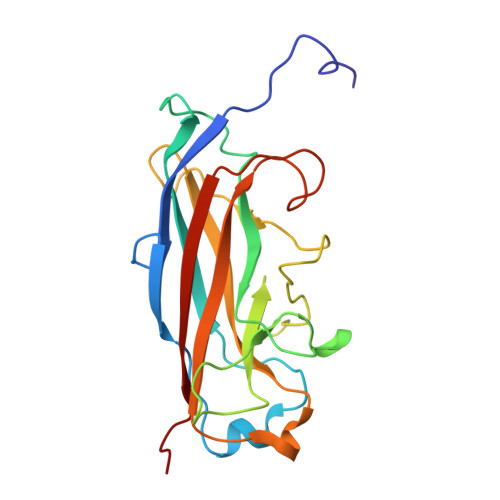
Biological and structural characterization of the Type 3 fimbrial subunit MrkA from Klebsiella pneumoniae.
Monaci, V., Oldrini, D., Gasperini, G., Banci, L., Cantini, F., Micoli, F.(2025) Protein Sci 34: e70343-e70343
- PubMed: 41118248 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.70343
- PubMed Abstract:
Klebsiella pneumoniae is a Gram-negative opportunistic pathogen responsible for a wide range of community-associated and hospital-acquired infections and a major cause of neonatal sepsis in low- and middle-income countries. The pathogen's surface fimbriae, particularly the Type 3 fimbriae, are critical for bacterial adhesion, biofilm formation, and host defense evasion. MrkA, the pathogen's major Type 3 fimbrial subunit, has a structural function in fimbrial assembly, but its three-dimensional structure remains to be fully elucidated. In this study, we utilized solution-state nuclear magnetic resonance spectroscopy to elucidate the structure of MrkA, leveraging previously reported chemical shift assignments of a designed self-complementing monomeric protein. Additionally, we confirmed the ability of monoclonal antibodies, capable of recognizing MrkA oligomers on wild-type Klebsiella bacteria, to bind the recombinant MrkA protein. Our findings contribute to the evaluation of MrkA as a potential target in vaccine development against Klebsiella pneumoniae infections.
- Magnetic Resonance Center - CERM, University of Florence, Florence, Italy.
Organizational Affiliation: