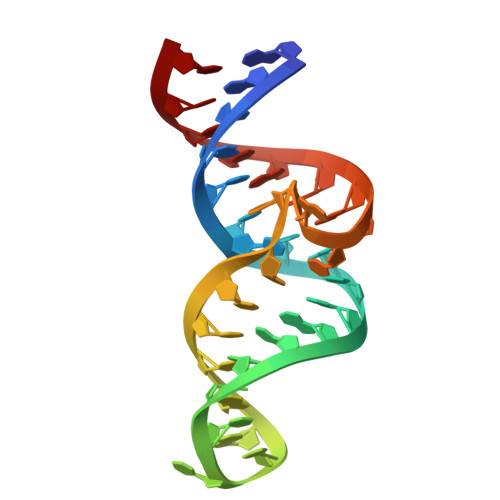
Structural basis for ligand recognition in the tobramycin riboswitch.
Duchardt-Ferner, E., Kraus, L., Limouchi, A., Suess, B., Wohnert, J.(2025) Nucleic Acids Res 53
- PubMed: 40902004 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkaf817
- Primary Citation Related Structures:
9HRO - PubMed Abstract:
Recently, a novel tobramycin-responsive riboswitch was developed by a combination of Capture-SELEX and in vivo screening. This riboswitch regulates translation initiation in eukaryotes with a high dynamic range and remarkable ligand affinity and selectivity. Its secondary structure differs from all previously described aminoglycoside-binding RNA motifs, suggesting a novel mode of ligand recognition. To provide a structural basis for the remarkable regulatory efficiency and ligand selectivity of this riboswitch, we investigated its structure in complex with its cognate ligand tobramycin by high-resolution solution nuclear magnetic resonance spectroscopy. The structure of the complex reveals a novel structural organization for an aminoglycoside binding motif with a unique pattern of intermolecular hydrogen bonds and electrostatic interactions between the RNA and functional groups of all three rings of the ligand. In contrast to other aminoglycoside binding motifs, ligand binding of the tobramycin riboswitch is coupled with the formation of an extensive network of noncanonical RNA-RNA interactions, rationalizing the high ligand affinity of this small hairpin RNA. Comparison with the free form of the RNA shows that the latter is much less compact, lacking many RNA-RNA interactions, in particular in the bulge regions, thereby immediately providing a rationale for the exceptional switching efficiency of this synthetic riboswitch.
- Institute for Molecular Biosciences and Center of Magnetic Resonance (BMRZ), Goethe-University Frankfurt, Max-von-Laue Straße 9, 60438 Frankfurt, Germany.
Organizational Affiliation: