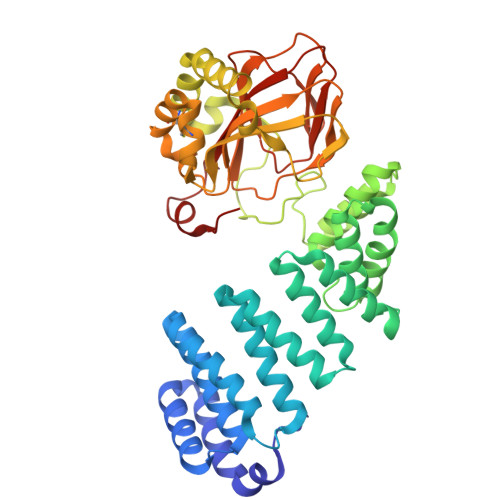
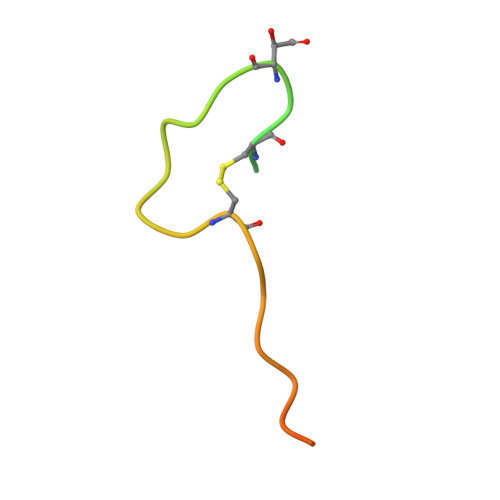
Structural basis of the promiscuity of the unusual Fe(II) and 2-oxoglutarate dependent human aspartate/asparagine-beta-hydroxylase.
de Munnik, M., Brasnett, A., Zhou, T., Myers, W., Wang, Y., Chatterjee, K., Tumber, A., Marshall, S.A., Simon, P.S., Aller, P., Shilova, A., Axford, D., Makita, H., Paley, D.W., Tiwari, V., Stead, A.T., Dehe, S., Sanchez, H., Rosenberg, D.J., Alonso-Mori, R., Bhowmick, A., Yano, J., Yachandra, V.K., Park, J., Park, S., Orville, A.M., Brewitz, L., Kern, J.F., Schofield, C.J., Rabe, P.(2026) Nat Commun
- PubMed: 41741441 Search on PubMed
- DOI: https://doi.org/10.1038/s41467-026-69425-w
- Primary Citation Related Structures:
9FVU, 9FVV, 9FVW, 9FVX, 9FVY, 9FVZ, 9FW0, 9HNZ, 9HO0, 9HO1, 9HO2, 9HO3 - PubMed Abstract:
Protein-hydroxylation catalysed by Fe(II) and 2-oxoglutarate (2OG) dependent oxygenases is an important regulatory mechanism in human biology. Such oxygenases typically coordinate their Fe(II) cofactor via a conserved triad of an aspartate- or glutamate- and two histidine-residues. By contrast, aspartate/asparagine β-hydroxylase (AspH), which catalyses asparagine/aspartate-residue oxidation in epidermal growth factor-like domains (EGFDs), has only two histidine-residues (H679, H725), with a water occupying the site normally occupied by an aspartate- or glutamate-residue. We describe mechanistic studies with catalytically active AspH crystals. Turnover studies with single crystals under cryogenic conditions give (3 R)-hydroxylated EGFDs with the product alcohol coordinating Fe(II) trans to H725. Time-resolved serial crystallography of microcrystals using an acoustic droplet ejection system, coupled to X-ray emission analyses, demonstrate turnover within 1.5 s, giving a product complex in which Fe(II) is regenerated. Solution and crystallographic studies with the O 2 surrogate nitric oxide imply O 2 binds to Fe(II) trans to H725. The additional Fe-chelating water is maintained throughout AspH catalysis and is not directly involved in substrate hydroxylation, because O 2 is the sole oxygen source in alcohol products, as shown by 18 O labelling studies. The results reveal how AspH accommodates both aspartate- and asparagine-substrates and will assist in efforts targeting AspH for cancer treatment.
- Chemistry Research Laboratory and the Ineos Oxford Institute for Antimicrobial Research, University of Oxford, Oxford, UK.
Organizational Affiliation: