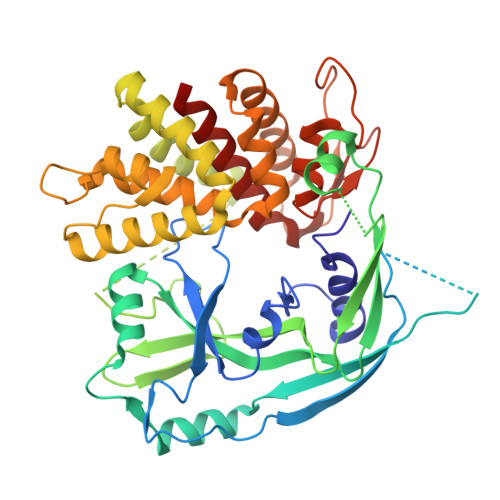
Improved structure of mouse gasdermin D: a new blueprint for structure-based drug design.
De Colibus, L., Ludzia, P., Biasutto, A., Pica, A., Hopper, J.T.S., Jazayeri, A., Durr, K.L.(2025) Acta Crystallogr F Struct Biol Commun 81: 408-415
- PubMed: 40906124 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X25007149
- Primary Citation Related Structures:
9HJP - PubMed Abstract:
Gasdermin D (GSDMD) is a protein that has gained significant attention in recent years due to its crucial role in inflammatory cell death, particularly pyroptosis. Pyroptosis is a highly inflammatory form of programmed cell death that is triggered by various microbial infections and sterile inflammatory stimuli. GSDMD acts as an executioner molecule in this process, leading to the release of pro-inflammatory cytokines and amplifying the immune response. Here, we present a higher resolution, significantly improved apo crystal structure of the deposited mouse structure model that will be beneficial for structure-based drug-design approaches towards this important pharmacological target.
- OMass Therapeutics, Building 4000, Chancellor Court, John Smith Drive, Oxford Business Park, ARC, Oxford OX4 2GX, United Kingdom.
Organizational Affiliation: