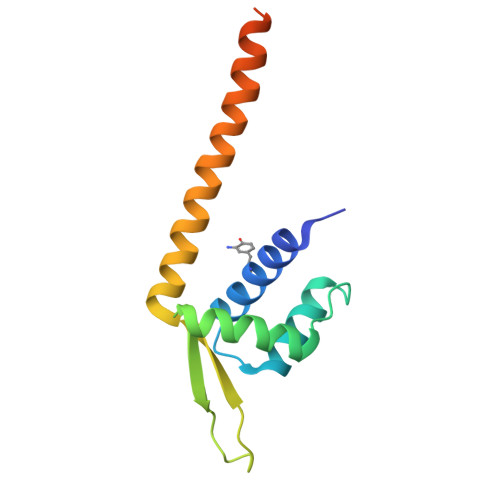
Genetically encoded 3-aminotyrosine as catalytic residue in a designer Friedel-Crafts alkylase.
Brouwer, B., Della-Felice, F., Thunnissen, A.W.H., Roelfes, G.(2025) Chem Sci 16: 8721-8728
- PubMed: 40276638 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/d5sc01055a
- Primary Citation Related Structures:
9H87, 9H88 - PubMed Abstract:
Genetic incorporation of noncanonical amino acids (ncAAs) harbouring catalytic side chains into proteins allows the creation of enzymes able to catalyse reactions that have no equivalent in nature. Here, we present for the first time the use of the ncAA 3-aminotyrosine (aY) as catalytic residue in a designer enzyme for iminium activation catalysis. Incorporation of aY into protein scaffold LmrR gave rise to an artificial Friedel-Crafts (FC) alkylase exhibiting complementary enantioselectivity to a previous FC-alkylase design using p -aminophenylalanine as catalytic residue in the same protein. The new FC-alkylase was optimized by directed evolution to afford a quadruple mutant that showed increased activity and excellent enantioselectivity (up to 95% ee). X-ray crystal structures of the parent and evolved designer enzymes suggest that the introduced mutations cause a narrowing of the active site and a reorientation of the catalytic -NH 2 group. Furthermore, the evolved FC-alkylase was applied in whole-cell catalysis, facilitated by the straightforward incorporation of aY. Our work demonstrates that aY is a valuable addition to the biochemists toolbox for creating artificial enzymes.
- Stratingh Institute for Chemistry, University of Groningen Nijenborgh 3 9747 AG Groningen The Netherlands j.g.roelfes@rug.nl.
Organizational Affiliation: