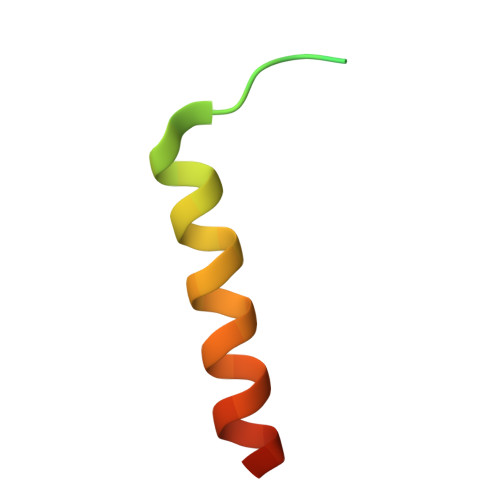
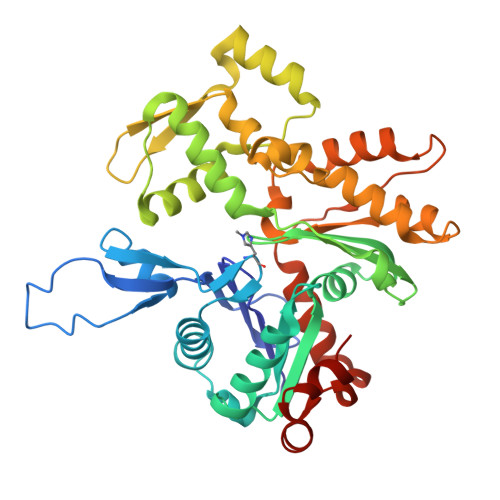
Structure of the F-tractin-F-actin complex.
Shatskiy, D., Sivan, A., Wedlich-Soldner, R., Belyy, A.(2025) J Cell Biol 224
- PubMed: 39928047 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1083/jcb.202409192
- Primary Citation Related Structures:
9GOB, 9HM9 - PubMed Abstract:
F-tractin is a peptide widely used to visualize the actin cytoskeleton in live eukaryotic cells but has been reported to impair cell migration and induce actin bundling at high expression levels. To elucidate these effects, we determined the cryo-EM structure of the F-tractin-F-actin complex, revealing that F-tractin consists of a flexible N-terminal region and an amphipathic C-terminal helix. The N-terminal part is dispensable for F-actin binding but responsible for the bundling effect. Based on these insights, we developed an optimized F-tractin, which eliminates the N-terminal region and minimizes bundling while retaining strong actin labeling. The C-terminal helix interacts with a hydrophobic pocket formed by two neighboring actin subunits, an interaction region shared by many actin-binding polypeptides, including the popular actin-binding probe Lifeact. Thus, rather than contrasting F-tractin and Lifeact, our data indicate that these peptides have analogous modes of interaction with F-actin. Our study dissects the structural elements of F-tractin and provides a foundation for developing future actin probes.
- Membrane Enzymology Group, Groningen Institute of Biomolecular Sciences and Biotechnology (GBB), Faculty of Science and Engineering, University of Groningen , Groningen, The Netherlands.
Organizational Affiliation: