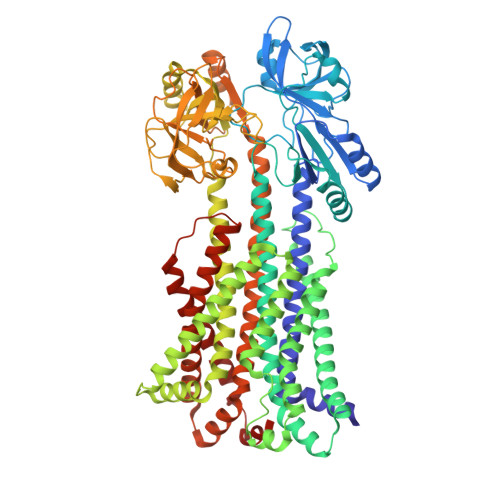
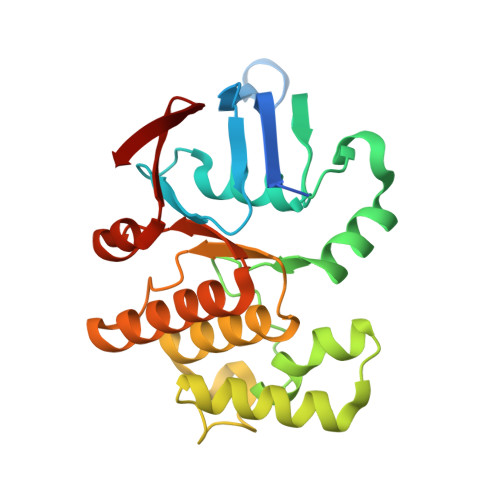
Structural characterization of the YbbAP-TesA ABC transporter identifies it as a lipid hydrolase complex that extracts hydrophobic compounds from the bacterial inner membrane.
McAndrew, M.B.L., Cook, J., Gill, A., Sahoo, K., Thomas, C., Stansfeld, P.J., Crow, A.(2025) PLoS Biol 23: e3003427-e3003427
- PubMed: 41289308 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pbio.3003427
- Primary Citation Related Structures:
9GE6, 9GE7, 9GE8 - PubMed Abstract:
Type VII ABC transporters are ATP-powered membrane protein complexes that drive key biological processes in the bacterial cell envelope. In Escherichia coli, three of the four Type VII ABC systems have been extensively characterized, including: the FtsEX-EnvC cell division complex, the LolCDE-LolA lipoprotein trafficking machinery, and the MacAB-TolC efflux pump. Here we describe a fourth E. coli Type VII ABC system, YbbAP-TesA, which combines a Type VII ABC transporter with a multifunctional hydrolytic enzyme. Structures of the complete YbbAP-TesA complex, and of YbbAP with and without bound ATP analogues, capture implied long-range transmembrane conformational changes that are the hallmark of this ABC superfamily's mechanotransmission mechanism. We further show that YbbAP-TesA can hydrolyze a variety of ester and thioester substrates and experimentally confirm a constellation of active site residues in TesA. Our data suggests YbbAP has a role in extracting hydrophobic molecules from the inner membrane and presenting these to TesA for hydrolysis. The work extends collective knowledge of the remarkable diversity of the ABC superfamily and establishes a new function for Type VII ABC transporters in bacterial cells.
- School of Life Sciences, University of Warwick, Coventry, United Kingdom.
Organizational Affiliation: