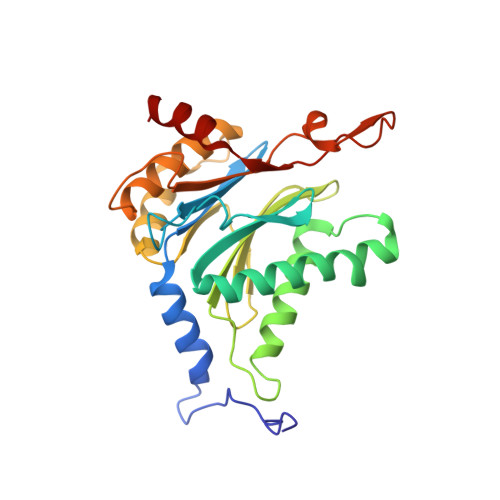
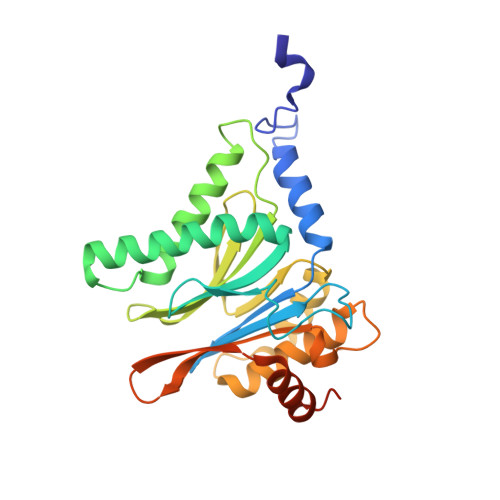
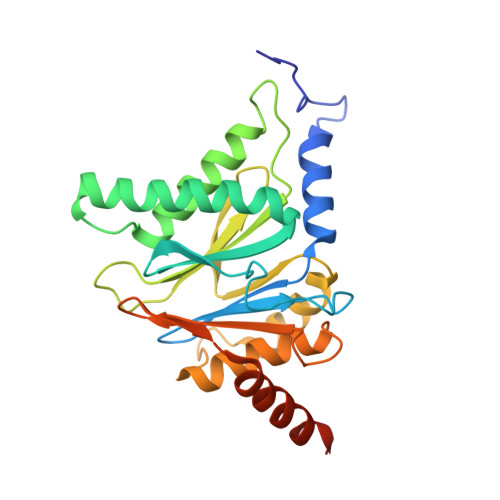
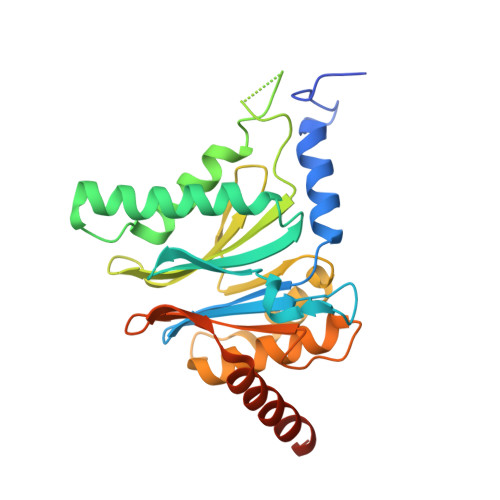
Structure-Based Design of Pan-Selective Peptide Epoxyketones for the Three Human Immunoproteasome Active Sites.
Dekker, P.M., Huber, E.M., Maurits, E., Wang, X., Heinemeyer, W., Florea, B.I., Groll, M., Overkleeft, H.S.(2026) J Med Chem 69: 1247-1263
- PubMed: 41504611 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.5c02639
- Primary Citation Related Structures:
9FRW, 9FST, 9FSU, 9FSV, 9FT0, 9FT1 - PubMed Abstract:
The proteasome inhibitors bortezomib, carfilzomib, and ixazomib all act by inhibiting multiple active sites of both constitutive proteasomes and immunoproteasomes. These clinical anticancer drugs are effective, but also display side effects, and evidence is amassing that their toxicity arises from constitutive proteasome inhibition. In this work, we describe the structure-guided discovery of a new class of pan-immunoproteasome-selective inhibitors. We identified the peptide epoxyketone BocPip-Ser ( 8 ), which targets all three human immunoproteasome active sites potently and with excellent selectivity over constitutive proteasome active sites (IC 50 values for i-subunits ≤ 0.92 μM; IC 50 ratio β1c/β1i: 13, β2c/β2i: 14, β5c/β5i: 18; Table 1 and Figure 3). We propose compound 8 (BocPip-Ser), which is of a similar size and general properties as carfilzomib, as a lead compound for the development of improved drugs targeting hematological cancers, and possibly also autoimmune diseases, driven by immunoproteasome but not constitutive proteasome activities.
- Leiden Institute of Chemistry, Leiden University, Einsteinweg 55, 2333 CC Leiden, The Netherlands.
Organizational Affiliation: