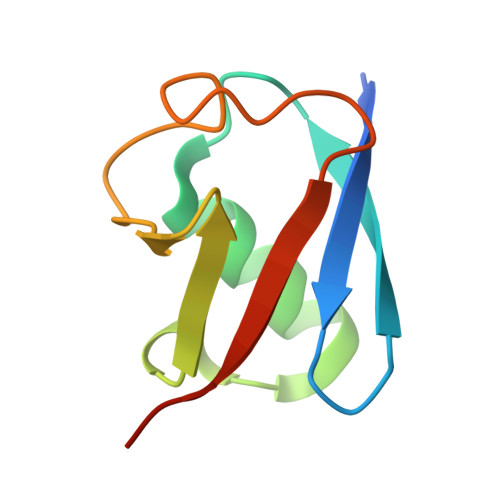
An engineered ubiquitin binding coiled coil peptide.
Vosbein, P., Vergara, P.P., Huang, D.T., Thomson, A.R.(2024) Chem Sci 15: 15776-15782
- PubMed: 39268210 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/d4sc04204b
- Primary Citation Related Structures:
9FJ3, 9FJ4 - PubMed Abstract:
Recognition of ubiquitin (Ub) is often mediated by small Ub binding domains such as the Ubiquitin Interacting Motif (UIM). Most Ub binding events are low affinity interactions, and designing stronger binders for Ub can be challenging. We here report the design of a short crosslinked coiled coil (CC) which is conformationally and chemically stable, and which can act as a scaffold to present the key binding residues from known UIM sequences. Doing so gives rise to a hybrid CC peptide that reconciles the important features of both UIM and CC sequences. We show by fluorescence polarization assays that this crosslinked 'CC-UIM' peptide exhibits enhanced binding to Ub compared to the original UIM sequence. Furthermore, we report a crystal structure of this peptide in complex with Ub. These studies show that preorganization of a small number of important binding residues onto a stable helical scaffold can be a successful strategy for binder design.
- School of Chemistry, University of Glasgow Glasgow G12 8QQ UK Drew.Thomson@glasgow.ac.uk.
Organizational Affiliation: