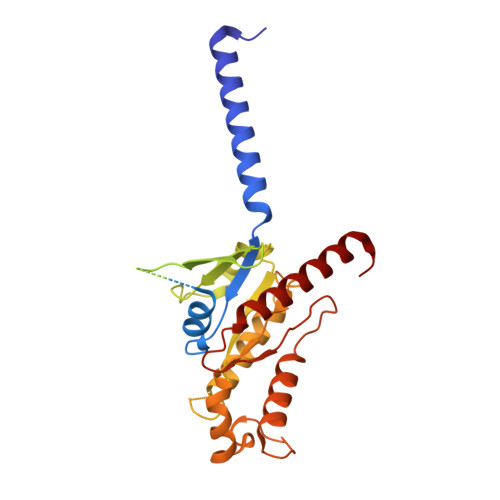
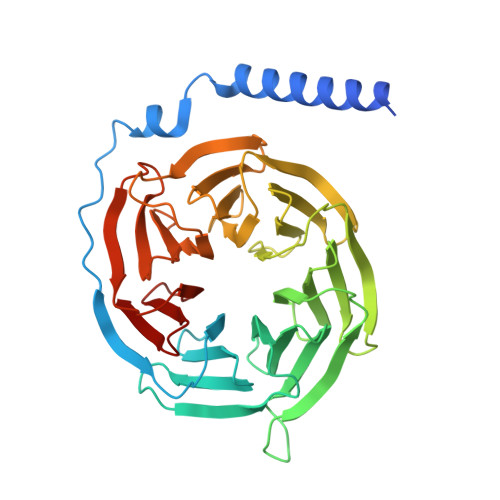
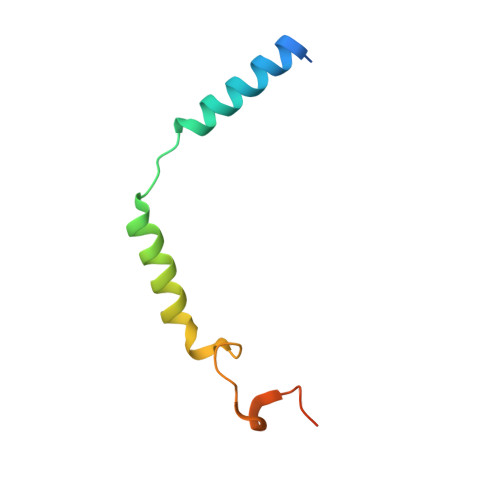
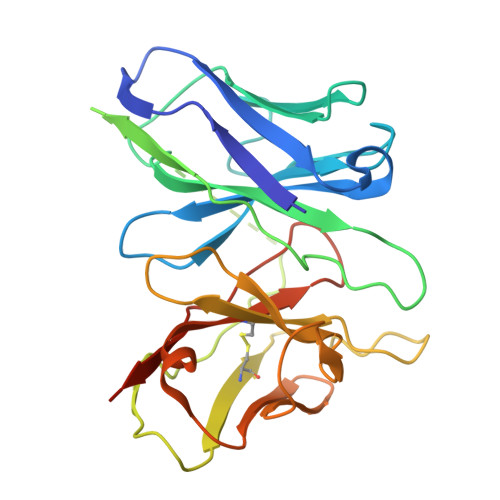
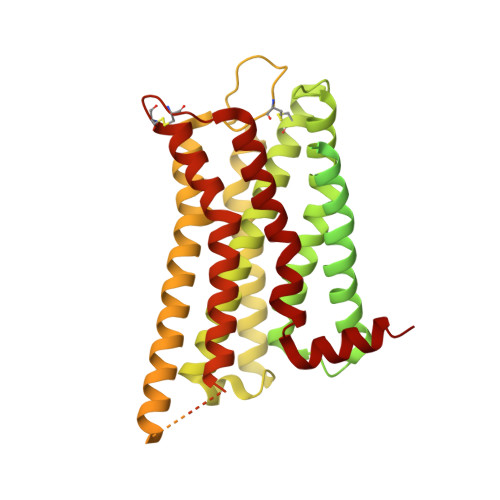
A bitopic agonist bound to the dopamine 3 receptor reveals a selectivity site.
Arroyo-Urea, S., Nazarova, A.L., Carrion-Antoli, A., Bonifazi, A., Battiti, F.O., Lam, J.H., Newman, A.H., Katritch, V., Garcia-Nafria, J.(2024) Nat Commun 15: 7759-7759
- PubMed: 39237617 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-51993-4
- Primary Citation Related Structures:
9F33, 9F34 - PubMed Abstract:
Although aminergic GPCRs are the target for ~25% of approved drugs, developing subtype selective drugs is a major challenge due to the high sequence conservation at their orthosteric binding site. Bitopic ligands are covalently joined orthosteric and allosteric pharmacophores with the potential to boost receptor selectivity and improve current medications by reducing off-target side effects. However, the lack of structural information on their binding mode impedes rational design. Here we determine the cryo-EM structure of the hD 3 R:Gα O βγ complex bound to the D 3 R selective bitopic agonist FOB02-04A. Structural, functional and computational analyses provide insights into its binding mode and point to a new TM2-ECL1-TM1 region, which requires the N-terminal ordering of TM1, as a major determinant of subtype selectivity in aminergic GPCRs. This region is underexploited in drug development, expands the established secondary binding pocket in aminergic GPCRs and could potentially be used to design novel and subtype selective drugs.
- Institute for Biocomputation and Physics of Complex Systems (BIFI), University of Zaragoza, Zaragoza, Spain.
Organizational Affiliation: