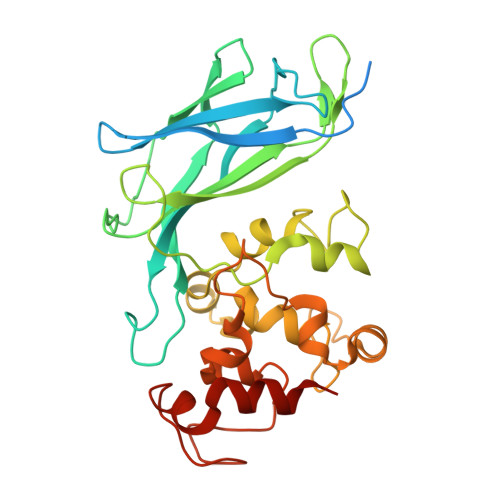
Extracellular catalysis of environmental substrates by Shewanella oneidensis MR-1 occurs via active sites on the C-terminal domains of MtrC.
Morales-Florez, A., Lockwood, C.W.J., Nash, B.W., Edwards, M.J., van Wonderen, J.H., Sachdeva, A., Butt, J.N., Clarke, T.A.(2025) Protein Sci 34: e70243-e70243
- PubMed: 40725986 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.70243
- Primary Citation Related Structures:
9EOV - PubMed Abstract:
The Gram-negative Shewanellaceae family is well known for its ability to transfer catabolically derived electrons to extracellular terminal electron acceptors through electron conduits that permeate the outer membrane. The primary conduit is MtrCAB, a trimeric porin-cytochrome complex that contains the cell surface exposed decaheme cytochrome MtrC. This donates electrons to extracellular substrates, including OmcA, soluble metals, organic electron shuttles, and insoluble metal oxides. However, it is not clear whether this broad substrate specificity requires specific sites for binding and reduction, or whether reduction occurs through non-specific interactions near exposed hemes on the cytochrome surface. Shewanella oneidensis MtrC is composed of four domains, with the hemes closely packed and distributed evenly between domains II and IV. The domains are arranged to allow electron transport across the cytochrome via interdomain electron transfer, but the significance of this conserved feature is not understood. Here we use site-directed mutagenesis to generate an MtrC variant that is comprised only of domains I and II (MtrC DI,II ). The properties of this MtrC DI,II are effectively identical to domains I and II of full-length MtrC. Whole-cell assays revealed that S. oneidensis cells replacing full-length MtrC with MtrC DI,II had significantly lower rates of OmcA, flavin mononucleotide, and Fe(III) citrate reduction. Our results demonstrate that MtrC domains III and IV contain sites for association of specific substrates, enabling the reduction of extracellular electron acceptors in S. oneidensis.
- School of Biological Sciences, University of East Anglia, Norwich, UK.
Organizational Affiliation: