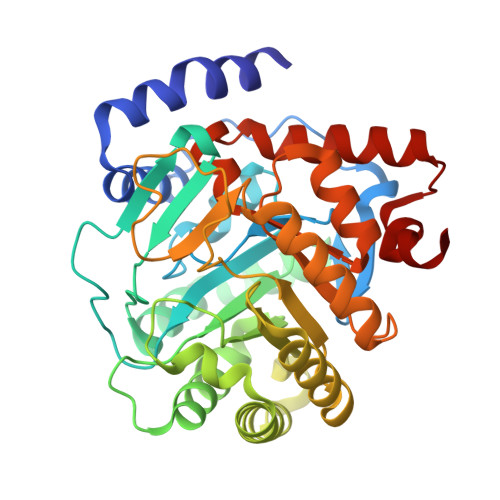
Crystallographic Structure of Human Dihydroorotate Dehydrogenase in Complex with the Natural Product Inhibitor Lapachol.
Purificacao, A.D., Benz, L.S., Lima Silva, W.J., Emery, F.S., Andrade, C.H., Weiss, M.S., Nonato, M.C.(2025) ACS Omega 10: 29087-29097
- PubMed: 40686985 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsomega.5c01536
- Primary Citation Related Structures:
9EG9 - PubMed Abstract:
Dihydroorotate dehydrogenase (DHODH) is a key enzyme in the pyrimidine biosynthesis pathway, playing a critical role in cellular processes and offering therapeutic potential for antiviral, antineoplastic, and autoimmune treatments. Human DHODH ( Hs DHODH) utilizes ubiquinone as a second substrate, positioning its quinone-binding site as a promising target for inhibitor development. Lapachol, a natural naphthoquinone, has gained prominence as a valuable natural product for the discovery of novel therapeutic agents, thanks to its wide range of biological activities. In this study, we present the first crystal structure of Hs DHODH in complex with lapachol, providing valuable insights into the interactions between this natural product and the enzyme. The structure reveals key binding interactions that mediate lapachol's affinity for Hs DHODH and validates previously proposed computational models. Complementary molecular dynamics simulations further highlight the stability of the complex and the importance of water-mediated interactions in ligand binding. These findings enhance our understanding of how naphthoquinone derivatives, such as lapachol, interact with class 2 DHODHs, offering a foundation for the design of optimized inhibitors for therapeutic applications. By integration of structural and computational data, this study contributes to the rational design of novel Hs DHODH inhibitors, paving the way for future exploration of lapachol and its derivatives in drug discovery.
- Center for the Research and Advancement in Fragments and Molecular Targets (CRAFT), School of Pharmaceutical Sciences at Ribeirao Preto, University of São Paulo, Ribeirão Preto 14040-903, São Paulo, Brazil.
Organizational Affiliation: