DNA-Encoded Library Screen Identifies Novel Series of Respiratory Syncytial Virus Polymerase Inhibitors.
Carney, S.M., Grosse, S., Yin, Y., Tran, M.T., Kalin, J.H., Jacoby, E., Fung, A., Simmons, N., Xie, X., Bhaumik, A., Carbajo, R.J., Piassek, M., Miller, R., Hu, L., Lemmens, C., Lutter, F.H., Pieters, S., Rombouts, G., Vetrano, I., Oehlrich, D., Tomaso, S., Lozada, K., Garcia, M.O., Anson, B., De Bruyn, S., Smith-Monroy, C., Neefs, J.M., Conceicao-Neto, N., Kesteleyn, B., Fino, R., Stoops, B., van Vlijmen, H., Patrick, A.N., Yu, X., Wong, V., Krosky, D.J., Abeywickrema, P., Ortiz-Meoz, R.F., Mason, S.W., Jin, Z., Sharma, S., Jonckers, T.H.M.(2025) J Med Chem 68: 6407-6430
- PubMed: 40042938 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.4c02906
- Primary Citation Related Structures:
9ECV, 9ED2 - PubMed Abstract:
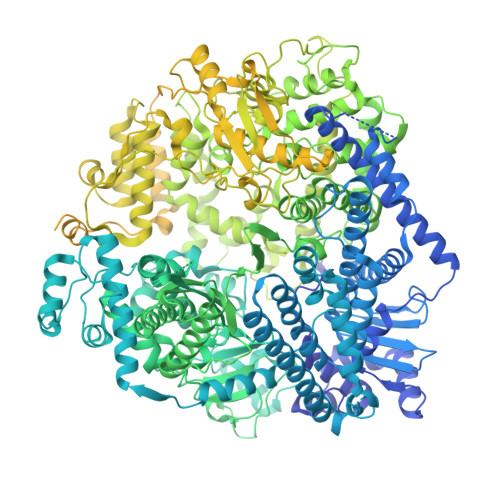
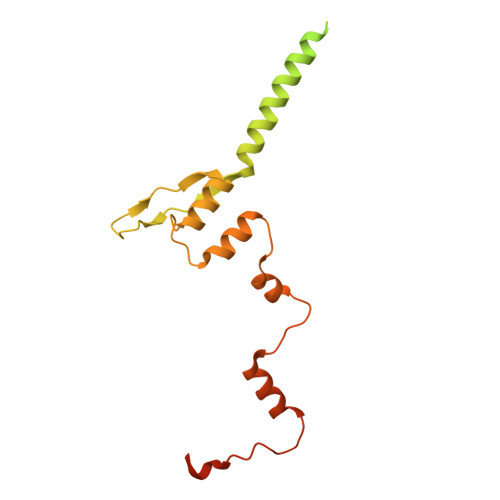
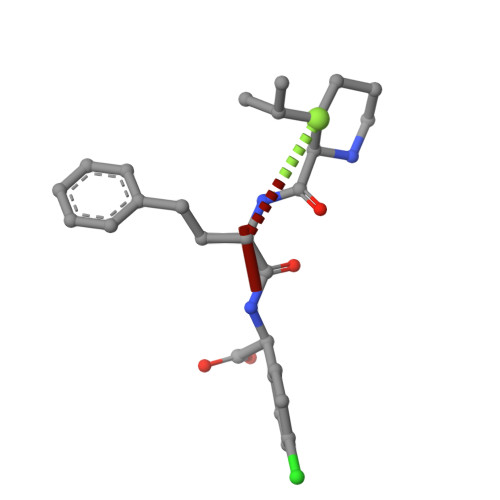
Respiratory syncytial virus (RSV) remains a public health burden due to unmet therapeutic needs. We recently reported the discovery of a non-nucleoside inhibitor of the RSV polymerase and characterized its binding to a novel pocket within the capping domain of the polymerase. Here, we describe our strategy to diversify the chemical matter targeting this site by screening our DNA-encoded chemical libraries, leading to the discovery of a novel and potent series of molecules that inhibits RSV polymerase's biochemical activity, as well as its viral replication in cells. Structural analysis via cryo-EM revealed novel contacts made within the capping domain binding pocket. By leveraging these structural insights for preliminary SAR exploration, we generated analogues for which potency and metabolic stability were improved more than 60- and 40-fold, respectively, over the initial hit. This work provides a path forward for further advanced SAR exploration and development of therapeutics against RSV.
- Janssen Research & Development, LLC, Johnson & Johnson Company, Spring House, Pennsylvania 19002, United States.
Organizational Affiliation: