Multimeric transcription factor BCL11A utilizes two zinc-finger tandem arrays to bind clustered short sequence motifs.
Horton, J.R., Yu, M., Zhou, J., Tran, M., Anakal, R.R., Lu, Y., Blumenthal, R.M., Zhang, X., Huang, Y., Zhang, X., Cheng, X.(2025) Nat Commun 16: 3672-3672
- PubMed: 40246927 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-58998-7
- Primary Citation Related Structures:
9E6R, 9E6S, 9E6T - PubMed Abstract:
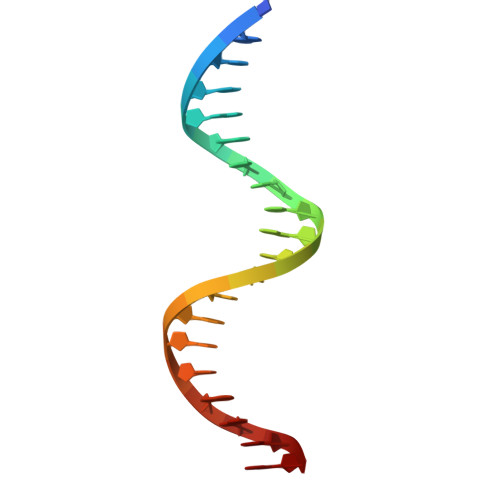
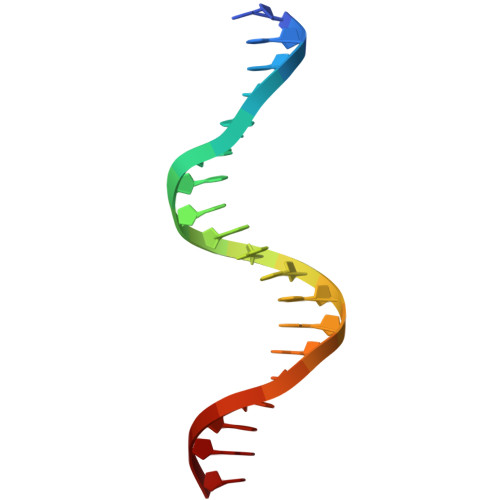
BCL11A, a transcription factor, is vital for hematopoiesis, including B and T cell maturation and the fetal-to-adult hemoglobin switch. Mutations in BCL11A are linked to neurodevelopmental disorders. BCL11A contains two DNA-binding zinc-finger arrays, low-affinity ZF2-3 and high-affinity ZF4-6, separated by a 300-amino-acid linker. ZF2-3 and ZF4-5 share 73% identity, including five out of six DNA base-interacting residues. These arrays bind similar short sequence motifs in clusters, with the linker enabling a broader binding span. Crystallographic structures of ZF4-6, in complex with oligonucleotides from the β-globin locus region, reveal DNA sequence recognition by residues Asn756 (ZF4), Lys784 and Arg787 (ZF5). A Lys784-to-Thr mutation, linked to a neurodevelopmental disorder with persistent fetal globin expression, reduces DNA binding over 10-fold but gains interaction with a variable base pair. BCL11A isoforms may form oligomers, enhancing chromatin occupancy and repressor functions by allowing multiple copies of both low- and high-affinity ZF arrays to bind DNA. These distinctive properties, apparently conserved among vertebrates, provide essential functional flexibility to this crucial regulator.
- Department of Epigenetics and Molecular Carcinogenesis, University of Texas MD Anderson Cancer Center, Houston, TX, 77030, USA.
Organizational Affiliation: