Crystal structure of the A/Viet Nam/1203/2004(H5N1) influenza virus hemagglutinin in complex with cyclic peptide iHA-100
Nguyen, T.K.Y., Wilson, I.A.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Hemagglutinin HA1 subunit | 323 | Influenza A virus | Mutation(s): 0 Gene Names: HA | 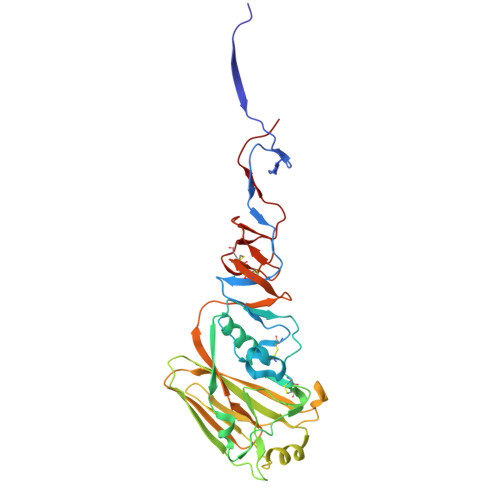 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A2P1MB68 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Hemagglutinin HA2 subunit | 176 | Influenza A virus | Mutation(s): 0 Gene Names: HA |  | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A2P1MB68 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| IHA-100 peptide | C [auth P] | 16 | Homo sapiens | Mutation(s): 0 |  |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 100.729 | α = 90 |
| b = 100.729 | β = 90 |
| c = 328.684 | γ = 120 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-2000 | data scaling |
| HKL-2000 | data reduction |
| PHENIX | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute Of Allergy and Infectious Diseases (NIH/NIAID) | United States | R56 AI117675 |