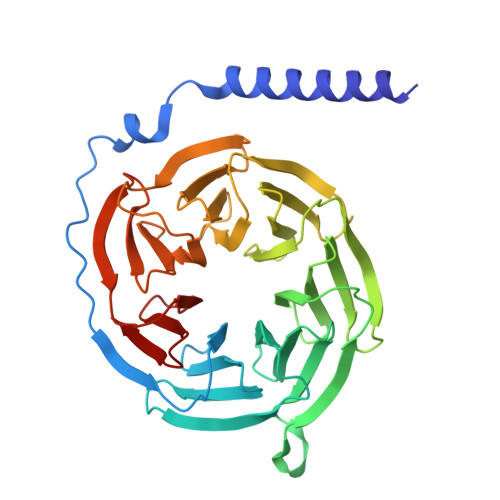
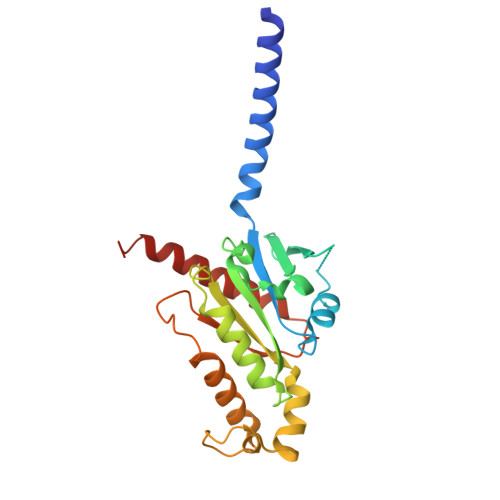
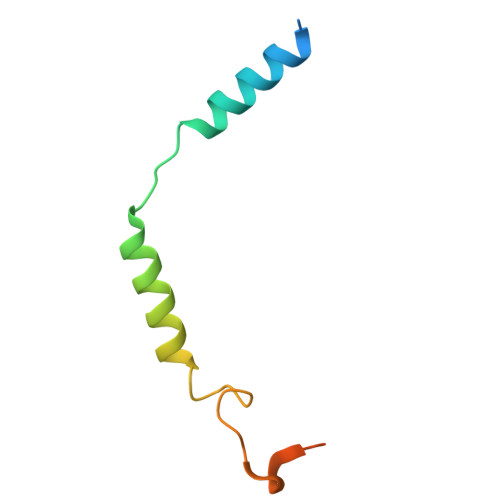
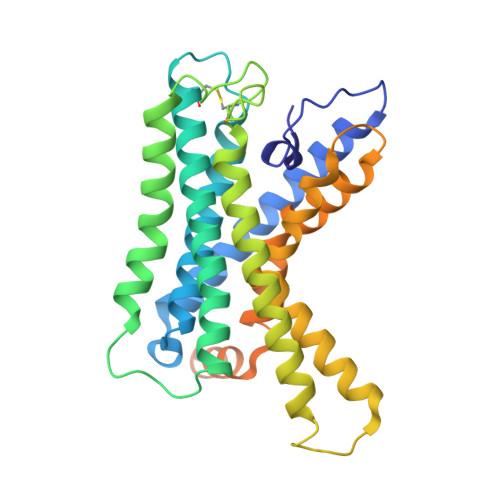
The adhesion GPCR ADGRL2 engages G alpha 13 to enable epidermal differentiation.
Yang, X., He, F., Lopez-Pajares, V., Porter, D.F., Garbett, K., Siprashvili, Z., Ducoli, L., Meyers, R.M., Reynolds, D.L., Lan Huong Bui, D., Hong, A., Nguyen, D.T., Jing, Y., Mondal, S., Ko, L., Tao, S., Singal, B., Sando, R., Skiniotis, G., Khavari, P.A.(2025) Proc Natl Acad Sci U S A 122: e2508436122-e2508436122
- PubMed: 41252157 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2508436122
- Primary Citation Related Structures:
9E51 - PubMed Abstract:
Homeostasis relies on signaling networks controlled by cell membrane receptors. Although G-protein-coupled receptors (GPCRs) are the largest family of transmembrane receptors, their specific roles in the epidermis are not fully understood. Dual CRISPR-Flow and single cell Perturb RNA-sequencing knockout screens of all epidermal GPCRs were thus performed, uncovering an essential requirement for adhesion GPCR ADGRL2 (latrophilin 2) in epidermal differentiation. Among potential downstream guanine nucleotide-binding G proteins, ADGRL2 selectively activated Gα13. Follow-up tissue knockouts verified that Gα13 is also required for epidermal differentiation. A cryoelectron microscopy structure in lipid nanodiscs showed that ADGRL2 engages with Gα13 at multiple interfaces, including via an interaction between ADGRL2 intracellular loop 3 and a Gα13-specific QQQ glutamine triplet sequence in its GTPase domain. In situ gene mutation of this interface sequence impaired epidermal differentiation, highlighting an essential new role for an ADGRL2-Gα13 axis in epidermal differentiation.
- Program in Epithelial Biology, Stanford University School of Medicine, Stanford, CA 94305.
Organizational Affiliation: