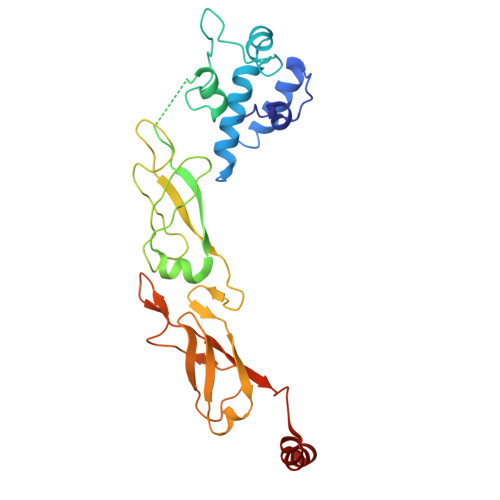
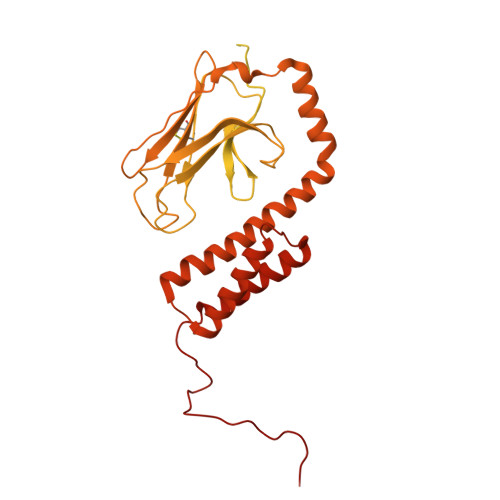
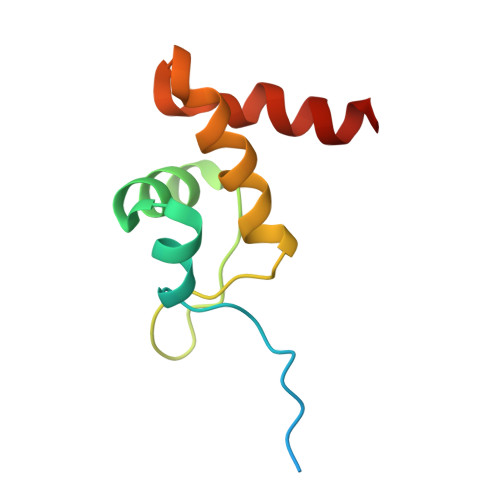
Mechanisms of assembly and function of the Hsp70-Hsp40 chaperone machinery.
Jiang, Y., Ibrahim, Z., Xia, Y., Clay, M., Myasnikov, A., Immadisetty, K., Xia, Z., Tang, L., Rossi, P., Ganguly, P., Liu, J., Miller, D., Che, M., Palacios, S.M., Kramer, G., Bukau, B., Kalodimos, C.G.(2025) Mol Cell 85: 4032-4046.e7
- PubMed: 41092901 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2025.09.023
- Primary Citation Related Structures:
8W1M, 9DVI, 9DYU, 9DYV - PubMed Abstract:
Hsp70 and Hsp40 molecular chaperones form a central machinery that remodels client proteins involved in numerous biological processes. Here, we integrated cryo-electron microscopy and nuclear magnetic resonance spectroscopy to determine the architecture of the full-length Hsp70-Hsp40 machinery. The structure of the complex in a physiologically inhibited state reveals distinct regulatory mechanisms. In the active state, the Hsp40 glycine-phenylalanine (G/F)-rich region acts as a pseudo-substrate for Hsp70, directly modulating refolding. This region also maintains Hsp40 in an autoinhibited state; upon binding to Hsp70, the inhibition is disrupted, exposing a cryptic client-binding site that enables client engagement and refolding. Transitions between these states are central to controlling refolding efficiency. Disrupting either the autoinhibited state or the G/F-Hsp70 interaction impairs function and elicits a compensatory heat shock response in cells. Our findings uncover the regulatory dynamics of a fundamental chaperone system, with broad implications for understanding protein homeostasis and the cellular response to stress.
- State Key Laboratory of Pharmaceutical Biotechnology, Department of Oncology, Nanjing Drum Tower Hospital, School of Life Sciences, Chemistry and Biomedicine Innovation Center (ChemBIC), Nanjing University, Nanjing, China; Department of Structural Biology, St. Jude Children's Research Hospital, Memphis, TN, USA.
Organizational Affiliation: