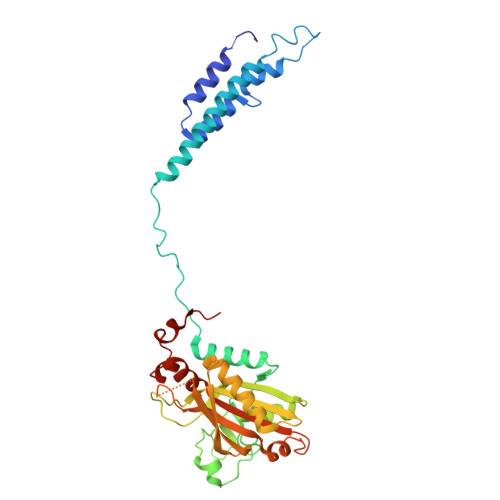
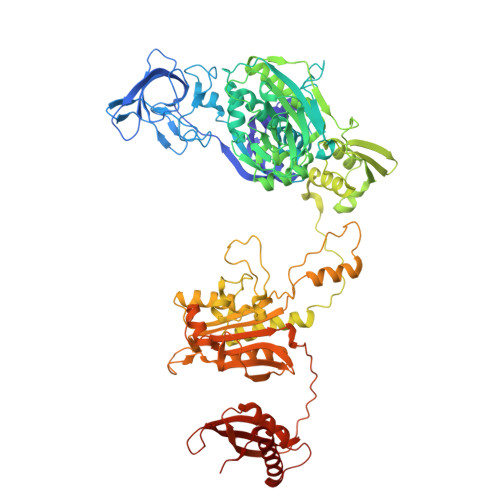
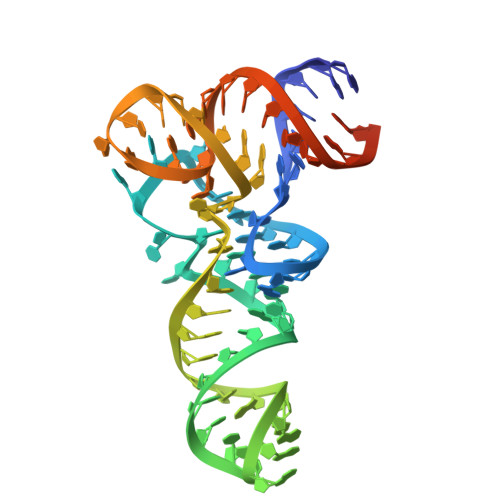
Different chemical scaffolds bind to L-phe site in Mycobacterium tuberculosis Phe-tRNA synthetase.
Gade, P., Chang, C., Pryde, D.S., Fletcher, D., Niven, S., Magalhaes, L.G., Robinson, D., Saini, J., Ibrahim, P.E.G.F., Forte, B., Wower, J., Bodkin, M.J., Baragana, B., Gilbert, I.H., Michalska, K., Joachimiak, A.(2025) Eur J Med Chem 287: 117335-117335
- PubMed: 39919438 Search on PubMed
- DOI: https://doi.org/10.1016/j.ejmech.2025.117335
- Primary Citation Related Structures:
9DRS, 9DRT, 9DRV, 9DSX, 9DTF - PubMed Abstract:
Tuberculosis (TB), caused by Mycobacterium tuberculosis (Mt), is one of the deadliest infectious diseases. The rise of multidrug-resistant strains represents a major public health threat, requiring new therapeutic options. Bacterial aminoacyl-tRNA synthetases (aaRS) have been shown to be highly promising drug targets, including for TB treatment. These enzymes play an essential role in translating the DNA gene code into protein sequence by attaching specific amino acid to their cognate tRNAs. They have multiple binding sites that can be targeted for inhibitor discovery: amino acid binding pocket, ATP binding pocket, tRNA binding site and an editing domain. Recently we reported several high-resolution structures of M. tuberculosis phenylalanyl-tRNA synthetase (MtPheRS) complexed with tRNA Phe and either L-Phe or a nonhydrolyzable phenylalanine adenylate analog. Here, using Nucleic Magnetic Resonance (NMR) and Surface Plasmon Resonance (SPR) we identified fragments that bind to MtPheRS and we determined crystal structures of their complexes with MtPheRS/tRNA Phe . All the binders interact with the L-Phe amino acid binding site. The analysis of interactions of the new compounds combined with adenylate analog structure provides insights for the rational design of anti-tuberculosis drugs. The 3' arm of the tRNA Phe in all the structures was disordered with exception of one complex with D-735 compound. In this structure the 3' CCA end of the acceptor stem is observed in the editing domain of MtPheRS providing insights regarding the post-transfer editing activity of class II aaRS.
- Center for Structural Biology of Infectious Diseases, Consortium for Advanced Science and Engineering, University of Chicago, Chicago, IL, 60667, USA; Structural Biology Center, X-ray Science Division, Argonne National Laboratory, Lemont, IL, 60439, USA.
Organizational Affiliation: