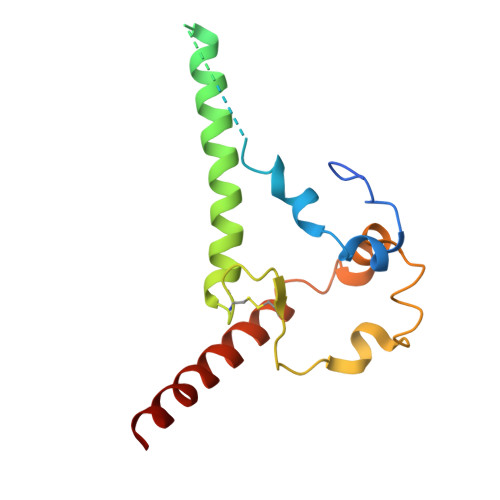
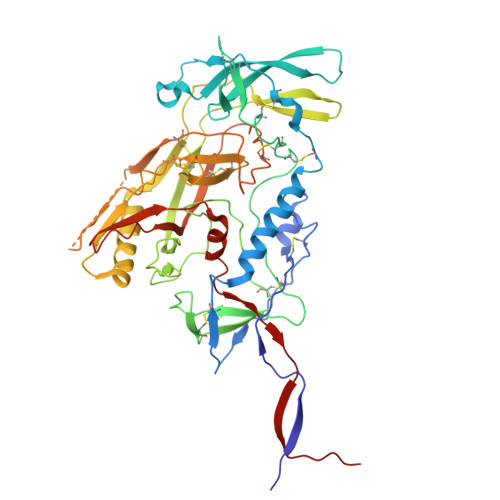
Transient glycan shield reduction induces CD4-binding site broadly neutralizing antibodies in SHIV-infected macaques.
Morris, D.J., Gorman, J., Zhou, T., Lora, J., Connell, A.J., Li, H., Liu, W., Roark, R.S., Campion, M.S., Carey, J.W., Habib, R., Li, Y., Martella, C.L., Park, Y., Singh, A., Sowers, K.J., Teng, I.T., Wang, S., Chohan, N., Ding, W., Lauer, C., Lewis, E., Mason, R.D., Rando, J.M., Peyton, L., Schramm, C.A., Wagh, K., Korber, B., Seaman, M.S., Douek, D.C., Haynes, B.F., Kulp, D.W., Roederer, M., Hahn, B.H., Kwong, P.D., Shaw, G.M.(2025) Cell Rep 44: 115848-115848
- PubMed: 40516049 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2025.115848
- Primary Citation Related Structures:
9DMB - PubMed Abstract:
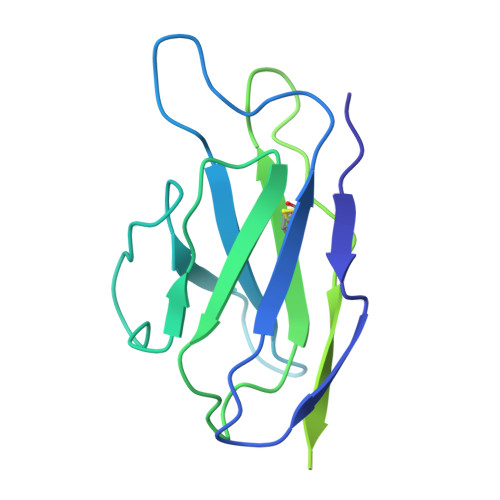
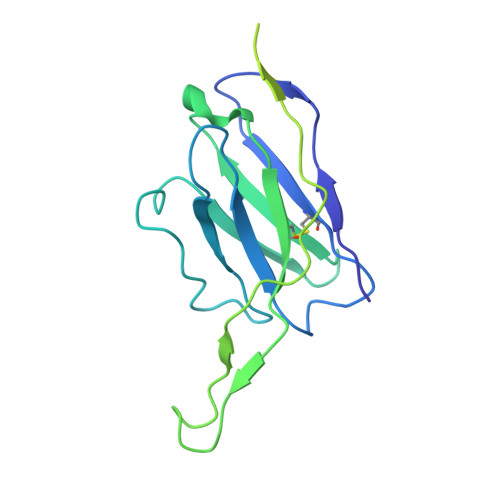
Broadly neutralizing antibodies (bNAbs) targeting the HIV-1 CD4-binding site (CD4bs) occur infrequently in macaques and humans and have not been reproducibly elicited in any outbred animal model. To address this challenge, we first isolated RHA10, an infection-induced rhesus bNAb with 51% breadth. The cryoelectron microscopy (cryo-EM) structure of RHA10 with the HIV-1 envelope (Env) resembled prototypic human CD4bs bNAbs with CDR-H3-dominated binding. Env-antibody co-evolution revealed transient elimination of two Env CD4bs-proximal glycans near the time of RHA10-lineage initiation, and these glycan-deficient Envs bound preferentially to early RHA10 intermediates, suggesting that glycan deletions in infecting SHIVs could induce CD4bs bNAbs. To test this hypothesis, we constructed SHIV.CH505 variants with CD4bs-proximal glycan deletions. Infection of 11 macaques resulted in accelerated CD4bs bNAb responses in 9 compared with 1 of 115 control macaques. Glycan hole-based immunofocusing coupled to Env-Ab co-evolution can consistently induce broad CD4bs responses in macaques and serve as a model for HIV vaccine design.
- Departments of Medicine and Microbiology, Perelman School of Medicine, University of Pennsylvania, Philadelphia, PA, USA.
Organizational Affiliation: