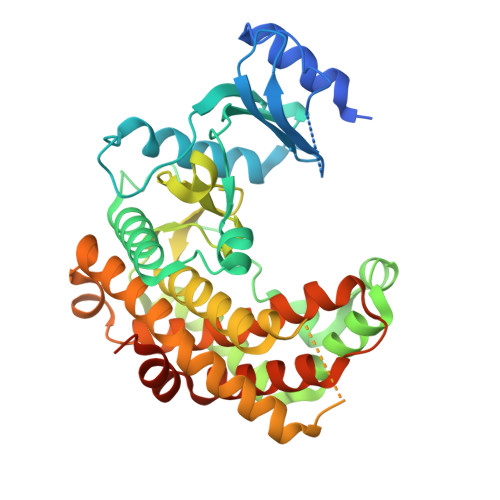
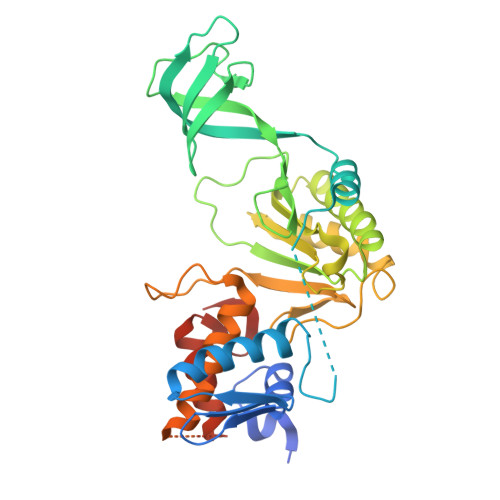
A Stable Dehydratase Complex Catalyzes the Formation of Dehydrated Amino Acids in a Class V Lanthipeptide.
Randall, G.T., Grant-Mackie, E.S., Chunkath, S., Williams, E.T., Middleditch, M.J., Tao, M., Harris, P.W.R., Brimble, M.A., Bashiri, G.(2024) ACS Chem Biol 19: 2548-2556
- PubMed: 39586055 Search on PubMed
- DOI: https://doi.org/10.1021/acschembio.4c00637
- Primary Citation Related Structures:
9DK1, 9DK2, 9DK3 - PubMed Abstract:
Lanthipeptides are ribosomally synthesized and post-translationally modified peptides that bear the characteristic lanthionine (Lan) or methyllanthionine (MeLan) thioether linkages. (Me)Lan moieties bestow lanthipeptides with robust stability and diverse antimicrobial, anticancer, and antiallodynic activities. Installation of (Me)Lan requires dehydration of serine and threonine residues to 2,3-dehydroalanine (Dha) and ( Z )-2,3-dehydrobutyrine (Dhb), respectively. LxmK and LxmY enzymes comprise the biosynthetic machinery of a newly discovered class V lanthipeptide, lexapeptide, and are proposed to catalyze the dehydration of serine and threonine residues in the precursor peptide. We demonstrate that LxmK and LxmY form a stable dehydratase complex to dehydrate precursor peptides. In addition, we present crystal structures of the LxmKY heterodimer, revealing structural and mechanistic features that enable iterative phosphorylation and elimination by the LxmKY complex. These findings provide molecular insights into class V lanthionine synthetases and lay the foundation for their applications as enzymatic tools in the biosynthesis of exquisitely modified peptides.
- School of Biological Sciences, The University of Auckland, Private Bag 92019, Auckland 1142, New Zealand.
Organizational Affiliation: