RTx-303, an Orally Bioavailable Pol theta Polymerase Inhibitor That Potentiates PARP Inhibitors in BRCA Mutant Tumors.
Chandramouly, G., Fried, W., Gordon, J., Ralph, D., Keuk, C., Kumari, S., Ramanjulu, M., Auerbacher, W., Minakhin, L., Tredinnick, T., Tiberi, B., Morton, G., Betsch, R., Cai, K.Q., Vekariya, U.M., Tyagi, M., Skorski, T., Karakashev, S., Johnson, N., Childers Jr., W.E., Chen, X.S., Pomerantz, R.T.(2025) J Med Chem 68: 22196-22215
- PubMed: 41124685 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.5c00551
- Primary Citation Related Structures:
9D6N, 9D6O - PubMed Abstract:
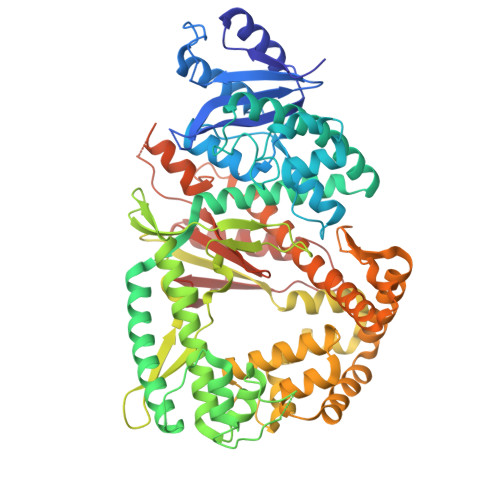
DNA polymerase θ (Polθ) is a polymerase-helicase fusion protein that is synthetically lethal with homologous recombination (HR) factors, such as BRCA1/2, and confers resistance to PARP inhibitors (PARPi) and other genotoxic cancer therapies. Previously developed Polθ polymerase (Polθ-pol) inhibitors (Polθi) exhibited limited pharmacological activity and metabolic stability, warranting the development of a Polθi with improved drug-like properties. Here, we developed RTx-303, a selective allosteric small-molecule Polθ-pol inhibitor that exhibits 5.1 nM IC 50 , 88% oral bioavailability, and a prolonged half-life along with its equipotent metabolite. X-ray crystallography highlights the development of a solvent-exposed side-chain that is essential for the optimal drug-like properties of RTx-303. Notably, RTx-303 exhibits significantly higher cellular potency than previously developed Polθ-pol inhibitors and strongly potentiates PARPi in BRCA1/2 mutant cells and patient-derived xenograft models. The superior potency, robust pharmacological activity, and high tolerability of RTx-303 warrant further development as a Polθ-pol inhibitor drug candidate.
- Department of Biochemistry and Molecular Biology, Sidney Kimmel Cancer Center, Thomas Jefferson University, Philadelphia, Pennsylvania 19107, United States.
Organizational Affiliation: