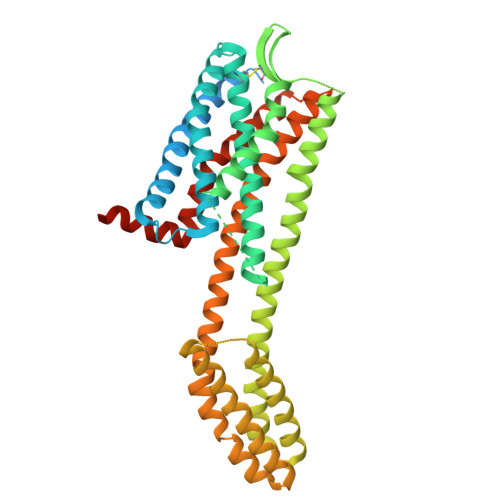
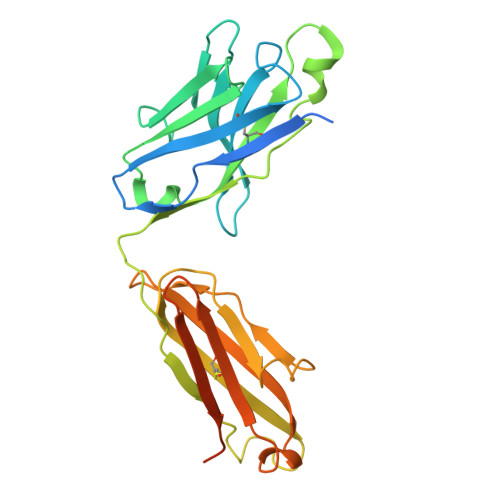
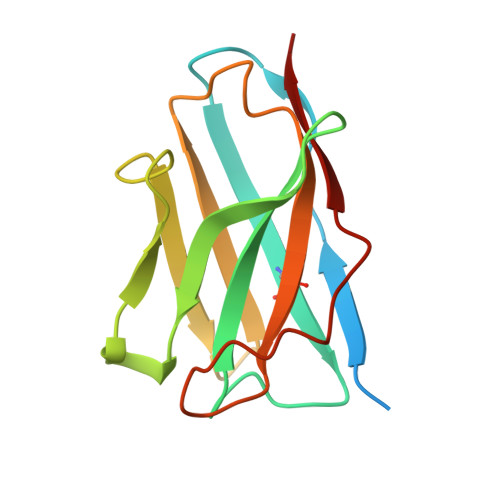
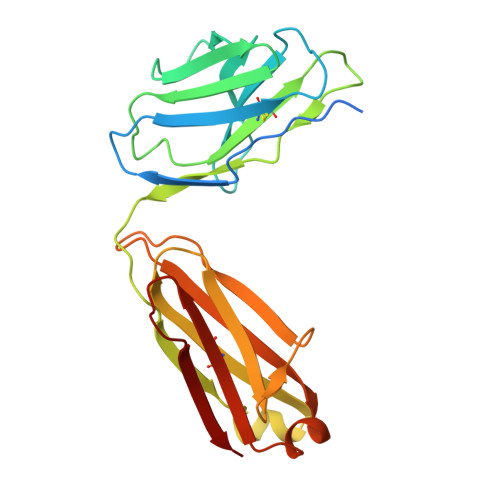
Structural basis for CCR6 modulation by allosteric antagonists.
Wasilko, D.J., Gerstenberger, B.S., Farley, K.A., Li, W., Alley, J., Schnute, M.E., Unwalla, R.J., Victorino, J., Crouse, K.K., Ding, R., Sahasrabudhe, P.V., Vincent, F., Frisbie, R.K., Dermenci, A., Flick, A., Choi, C., Chinigo, G., Mousseau, J.J., Trujillo, J.I., Nuhant, P., Mondal, P., Lombardo, V., Lamb, D., Hogan, B.J., Minhas, G.S., Segala, E., Oswald, C., Windsor, I.W., Han, S., Rappas, M., Cooke, R.M., Calabrese, M.F., Berstein, G., Thorarensen, A., Wu, H.(2024) Nat Commun 15: 7574-7574
- PubMed: 39217154 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-52045-7
- Primary Citation Related Structures:
9D3E, 9D3G - PubMed Abstract:
The CC chemokine receptor 6 (CCR6) is a potential target for chronic inflammatory diseases. Previously, we reported an active CCR6 structure in complex with its cognate chemokine CCL20, revealing the molecular basis of CCR6 activation. Here, we present two inactive CCR6 structures in ternary complexes with different allosteric antagonists, CCR6/SQA1/OXM1 and CCR6/SQA1/OXM2. The oxomorpholine analogues, OXM1 and OXM2 are highly selective CCR6 antagonists which bind to an extracellular pocket and disrupt the receptor activation network. An energetically favoured U-shaped conformation in solution that resembles the bound form is observed for the active analogues. SQA1 is a squaramide derivative with close-in analogues reported as antagonists of chemokine receptors including CCR6. SQA1 binds to an intracellular pocket which overlaps with the G protein site, stabilizing a closed pocket that is a hallmark of inactive GPCRs. Minimal communication between the two allosteric pockets is observed, in contrast to the prevalent allosteric cooperativity model of GPCRs. This work highlights the versatility of GPCR antagonism by small molecules, complementing previous knowledge of CCR6 activation, and sheds light on drug discovery targeting CCR6.
- Discovery Sciences, Medicine Design, Pfizer Inc., Groton, CT, USA.
Organizational Affiliation: