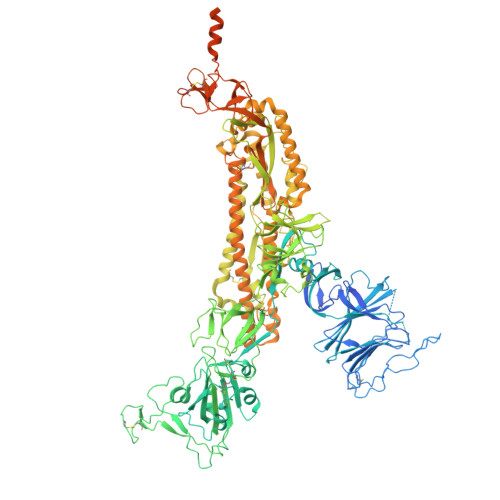
Production and cryo-electron microscopy structure of an internally tagged SARS-CoV-2 spike ecto-domain construct.
Singh, S., Liu, Y., Burke, M., Rayaprolu, V., Stein, S.E., Hasan, S.S.(2025) J Struct Biol X 11: 100123-100123
- PubMed: 40046771 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.yjsbx.2025.100123
- Primary Citation Related Structures:
9B0Y, 9B2V, 9CSS, 9CT2, 9CVH, 9CXE, 9N2L - PubMed Abstract:
The SARS-CoV-2 spike protein is synthesized in the endoplasmic reticulum of host cells, from where it undergoes export to the Golgi and the plasma membrane or retrieval from the Golgi to the endoplasmic reticulum. Elucidating the fundamental principles of this bidirectional secretion are pivotal to understanding virus assembly and designing the next generation of spike genetic vaccine with enhanced export properties. However, the widely used strategy of C-terminal affinity tagging of the spike cytosolic tail interferes with proper bidirectional trafficking. Hence, the structural and biophysical investigations of spike protein trafficking have been hindered by a lack of appropriate spike constructs. Here we describe a strategy for the internal tagging of the spike protein. Using sequence analyses and AlphaFold modeling, we identified a site down-stream of the signal sequence for the insertion of a twin-strep-tag, which facilitates purification of an ecto-domain construct from the extra-cellular medium of mammalian Expi293F cells. Mass spectrometry analyses show that the internal tag has minimal impact on N -glycan modifications, which are pivotal for spike-host interactions. Single particle cryo-electron microscopy reconstructions of the spike ecto-domain reveal conformational states compatible for ACE2 receptor interactions, further solidifying the feasibility of the internal tagging strategy. Collectively, these results present a substantial advance towards reagent development for the investigations of spike protein trafficking during coronavirus infection and genetic vaccination.
- Department of Biochemistry and Molecular Biology, University of Maryland School of Medicine, Baltimore MD 21201, USA.
Organizational Affiliation: