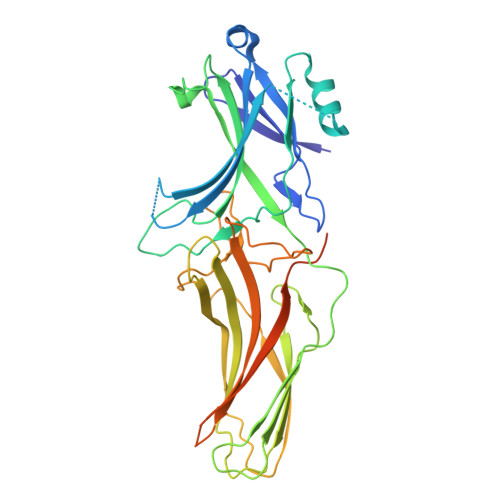
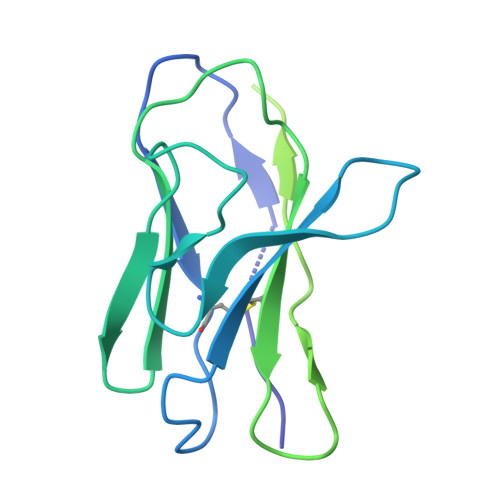
Mechanism of beta-arrestin 1 mediated Src activation via Src SH3 domain revealed by cryo-electron microscopy.
Pakharukova, N., Thomas, B.N., Bansia, H., Li, L., Bassford, D.K., Abzalimov, R.R., Kim, J., Kahsai, A.W., Pani, B., Xiao, K., Ochakovski, R., Liu, S., Zhang, X., Ahn, S., des Georges, A., Lefkowitz, R.J.(2026) Nat Commun 17
- PubMed: 41720803 Search on PubMed
- DOI: https://doi.org/10.1038/s41467-026-69884-1
- Primary Citation Related Structures:
9BT8, 9CX3, 9CX9 - PubMed Abstract:
Beta-arrestins (βarrs) are key regulators and transducers of G-protein coupled receptor signaling; however, little is known of how βarrs communicate with their downstream effectors. Here, we delineate structural mechanisms underlying βarr-mediated signal transduction. Using cryo-electron microscopy, we elucidate how βarr1 recruits and activates the non-receptor tyrosine kinase Src, a well-established signaling partner of βarrs. βarr1 engages Src SH3 through two distinct sites, each employing a different recognition mechanism: a polyproline motif in the N-domain and a non-proline-based interaction in the central crest region. At both sites βarr1 interacts with the aromatic surface of SH3, disrupting the autoinhibited conformation of Src and directly triggering its allosteric activation. This structural evidence establishes βarr1 as an active regulatory protein rather than a passive scaffold and suggests a potentially general mechanism for βarr-mediated signaling across diverse effectors.
- Department of Medicine, Duke University Medical Center, Durham, NC, 27710, USA.
Organizational Affiliation: