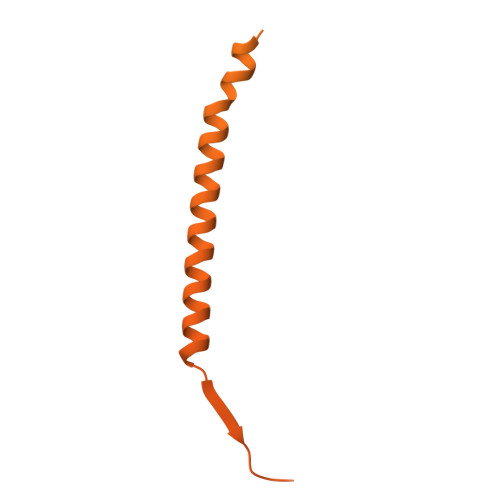
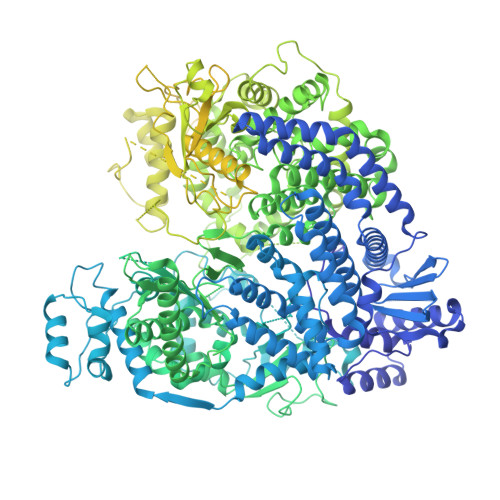
Cryo-EM structures of Nipah virus polymerases and high-throughput RdRp assay development enable anti-NiV drug discovery.
Chen, Z., Quirit Dudley, J., Deniston, C., Buffalo, C.Z., Patra, D., Cao, D., Hunt, J., Rohaim, A., Sengupta, D., Wen, L., Tsang, T., Xie, L., DiDonato, M., Spraggon, G., Clifton, M.C., Jarrousse, N., Straimer, J., Liang, B.(2025) Nat Commun 16: 6655-6655
- PubMed: 40683863 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-61764-4
- Primary Citation Related Structures:
9COK, 9MUW, 9MZH - PubMed Abstract:
Transcription and replication of the Nipah virus (NiV) are driven by the large protein (L) together with its essential co-factor phosphoprotein (P). L encodes all the viral enzymatic functions, including RNA-dependent RNA polymerase (RdRp) activity, while the tetrameric P is multi-modular. Here, we investigate the molecular mechanism of the NiV polymerase and build tools for anti-NiV drug discovery. We analyze and compare multiple cryo-EM structures of both full-length and truncated NiV polymerases from the Malaysia and Bangladesh strains. We identify two conserved loops in the polyribonucleotidyltransferase (PRNTase) domain of L and the binding between RdRp-PRNTase and CD domains. To further assess the mechanism of NiV polymerase activity, we establish a highly sensitive radioactive-labeled RNA synthesis assay and identify a back-priming activity in the NiV polymerase as well as a fluorescence and luminescent-based non-radioactive polymerase assay to enable high-throughput screening for L protein inhibitors. The combination of structural analysis and the development of both high-sensitive and high-throughput biochemical assays will enable the identification of new direct-acting antiviral candidates for treating highly pathogenic henipaviruses.
- Department of Biochemistry, Emory University School of Medicine, Atlanta, GA, USA.
Organizational Affiliation: