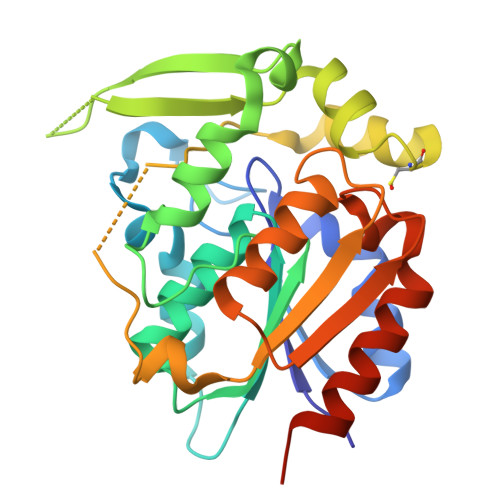
Crystal structures of 40- and 71-substitution variants of hydroxynitrile lyase from rubber tree.
Pierce, C.T., Tan, P., Greenberg, L.R., Walsh, M.E., Shi, K., Nguyen, A.H., Meixner, E.L., Sarak, S., Aihara, H., Evans 3rd, R.L., Kazlauskas, R.J.(2025) Acta Crystallogr D Struct Biol 81: 511-523
- PubMed: 40864494 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798325007065
- Primary Citation Related Structures:
8SNI, 9CLR - PubMed Abstract:
Hydroxynitrile lyase from Hevea brasiliensis (HbHNL) and the esterase SABP2 from Nicotiana tabacum share the α/β-hydrolase fold, a Ser-His-Asp catalytic triad and 44% sequence identity, yet catalyze different reactions. Prior studies showed that three active-site substitutions in HbHNL conferred weak esterase activity. To investigate how regions beyond the active site influence catalytic efficiency and active-site geometry, we engineered HbHNL variants with increasing numbers of substitutions to match SABP2. Variant HNL16 has all amino acids within 6.5 Å of the active site identical to SABP2, HNL40 those within 10 Å and HNL71 those within 14 Å. HNL16 exhibited poor esterase activity, whereas both HNL40 and HNL71 showed efficient esterase catalysis, demonstrating that residues beyond the immediate active site are critical for functional conversion. X-ray structures of HNL40 and HNL71 reveal a progressive shift in backbone positions toward those of SABP2, with r.m.s.d. values of 0.51 Å (HNL40) and 0.41 Å (HNL71) over the C α atoms, and even smaller r.m.s.d.s within the active-site region. Both HNL40 and HNL71 show a restored oxyanion hole and an additional tunnel connecting the active site to the protein surface. This work demonstrates the essential role of distant, indirectly acting residues to catalysis in α/β-hydrolase enzymes.
- Department of Biochemistry, Molecular Biology and Biophysics, University of Minnesota, Minneapolis, MN 55455, USA.
Organizational Affiliation: