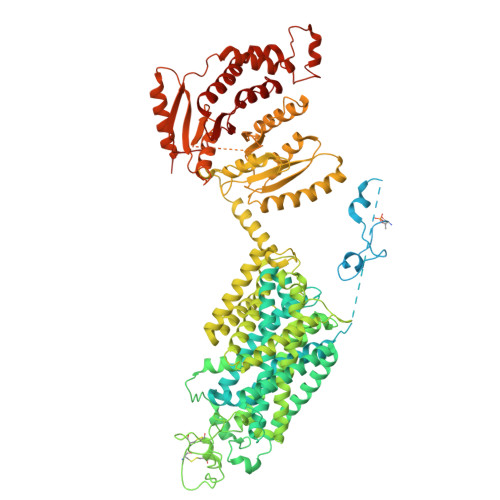
Structural basis for human NKCC1 inhibition by loop diuretic drugs.
Zhao, Y., Vidossich, P., Forbush, B., Ma, J., Rinehart, J., De Vivo, M., Cao, E.(2025) EMBO J 44: 1540-1562
- PubMed: 39875725 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s44318-025-00368-6
- Primary Citation Related Structures:
9C0E, 9C0G, 9C0H - PubMed Abstract:
Na + -K + -Cl - cotransporters functions as an anion importers, regulating trans-epithelial chloride secretion, cell volume, and renal salt reabsorption. Loop diuretics, including furosemide, bumetanide, and torsemide, antagonize both NKCC1 and NKCC2, and are first-line medicines for the treatment of edema and hypertension. NKCC1 activation by the molecular crowding sensing WNK kinases is critical if cells are to combat shrinkage during hypertonic stress; however, how phosphorylation accelerates NKCC1 ion transport remains unclear. Here, we present co-structures of phospho-activated NKCC1 bound with furosemide, bumetanide, or torsemide showing that furosemide and bumetanide utilize a carboxyl group to coordinate and co-occlude a K + , whereas torsemide encroaches and expels the K + from the site. We also found that an amino-terminal segment of NKCC1, once phosphorylated, interacts with the carboxyl-terminal domain, and together, they engage with intracellular ion exit and appear to be poised to facilitate rapid ion translocation. Together, these findings enhance our understanding of NKCC-mediated epithelial ion transport and the molecular mechanisms of its inhibition by loop diuretics.
- Department of Biochemistry, University of Utah School of Medicine, Salt Lake City, UT, 84112-5650, USA.
Organizational Affiliation: