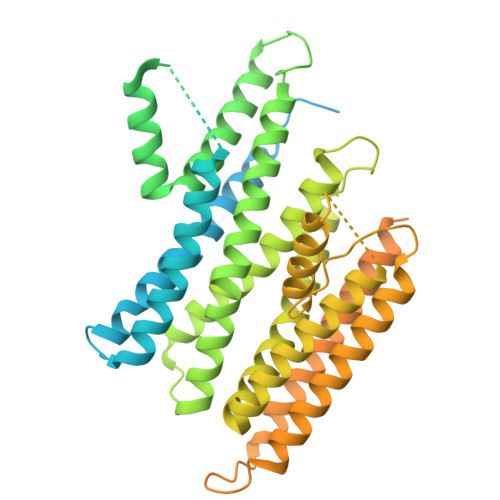
Structure and function of the human apoptotic scramblase Xkr4.
Chakraborty, S., Feng, Z., Lee, S., Alvarenga, O.E., Panda, A., Zhang, S., Bruni, R., Khelashvili, G., Gupta, K., Accardi, A.(2025) Nat Commun 16: 7317-7317
- PubMed: 40781244 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-62739-1
- Primary Citation Related Structures:
9BOJ - PubMed Abstract:
Phosphatidylserine externalization on the surface of dying cells is a key signal for their recognition and clearance by macrophages and is mediated by members of the X-Kell related (Xkr) protein family. Defective Xkr-mediated scrambling impairs clearance, leading to inflammation. It was proposed that activation of the Xkr4 apoptotic scramblase requires caspase cleavage, followed by dimerization and ligand binding. Here, using a combination of biochemical approaches we show that purified monomeric, full-length human Xkr4 (hXkr4) scrambles lipids. CryoEM imaging shows that hXkr4 adopts a novel conformation, where three conserved acidic residues create a negative electrostatic surface embedded in the membrane. Molecular dynamics simulations show this conformation induces membrane thinning, which could promote scrambling. Thinning is ablated or reduced in conditions where scrambling is abolished or reduced. Our work provides insights into the molecular mechanisms of hXkr4 scrambling and suggests the ability to thin membranes might be a general property of active scramblases.
- Department of Anesthesiology, Weill Cornell Medical College, New York, NY, 10027, USA.
Organizational Affiliation: