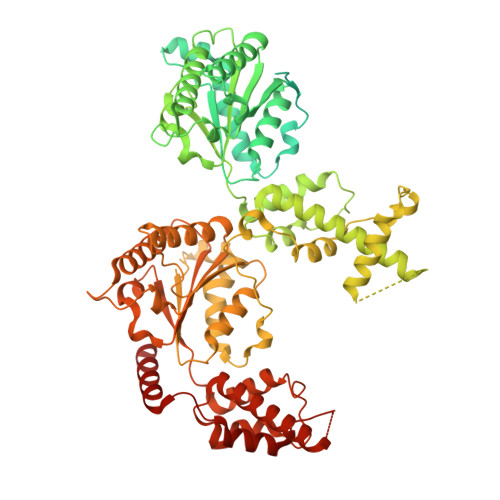
Dibenzothiazepinones are bioavailable inhibitors of NVL and cause p53-dependent apoptosis in cancer by blocking 60S ribosome assembly
Guo, H.H., Tao, Y., Cruz, V.E., Fang, M., Khivansara, V., Xie, S., Leach, A., Rivera-Cancel, G., Barrows, N., Williams, N., Aurora, A., Erzberger, J.P., De Brabander, J.K., Nijhawan, D.To be published.