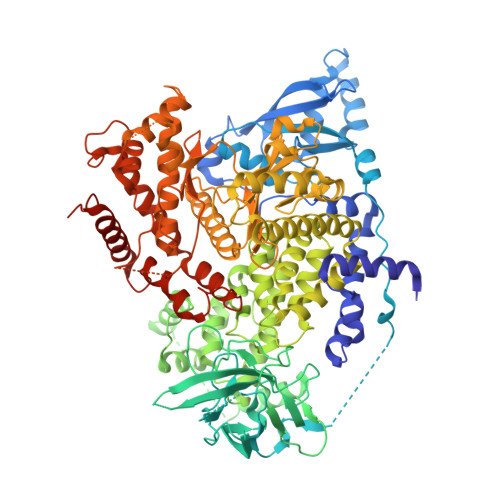
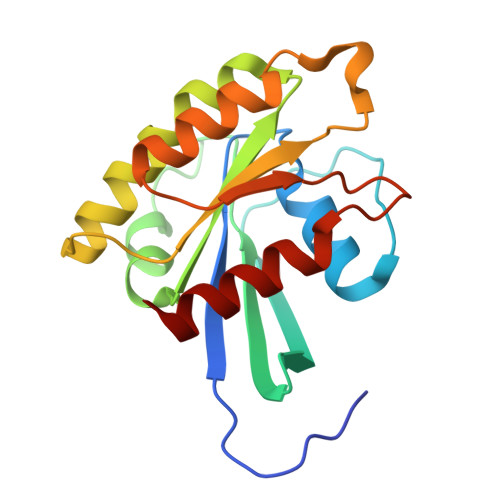
Structural insights into isoform-specific RAS-PI3K alpha interactions and the role of RAS in PI3K alpha activation.
Czyzyk, D., Yan, W., Messing, S., Gillette, W., Tsuji, T., Yamaguchi, M., Furuzono, S., Turner, D.M., Esposito, D., Nissley, D.V., McCormick, F., Simanshu, D.K.(2025) Nat Commun 16: 525-525
- PubMed: 39788953 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-55766-x
- Primary Citation Related Structures:
9B4Q, 9B4R, 9B4S, 9B4T, 9C15 - PubMed Abstract:
Mutations in RAS and PI3Kα are major drivers of human cancer. Their interaction plays a crucial role in activating PI3Kα and amplifying the PI3K-AKT-mTOR pathway. Disrupting RAS-PI3Kα interaction enhances survival in lung and skin cancer models and reduces tumor growth and angiogenesis, although the structural details of this interaction remain unclear. Here, we present structures of KRAS, RRAS2, and MRAS bound to the catalytic subunit (p110α) of PI3Kα, elucidating the interaction interfaces and local conformational changes upon complex formation. Structural and mutational analyses highlighted key residues in RAS and PI3Kα impacting binding affinity and revealed isoform-specific differences at the interaction interface in RAS and PI3K isoforms, providing a rationale for their differential affinities. Notably, in the RAS-p110α complex structures, RAS interaction with p110α is limited to the RAS-binding domain and does not involve the kinase domain. This study underscores the pivotal role of the RAS-PI3Kα interaction in PI3Kα activation and provides a blueprint for designing PI3Kα isoform-specific inhibitors to disrupt this interaction.
- NCI RAS Initiative, Cancer Research Technology Program, Frederick National Laboratory for Cancer Research, Frederick, MD, USA.
Organizational Affiliation: