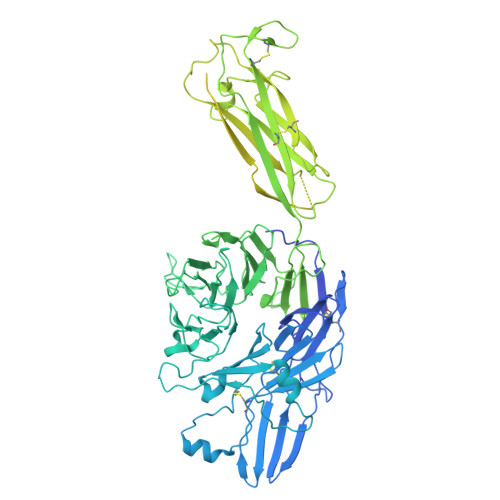
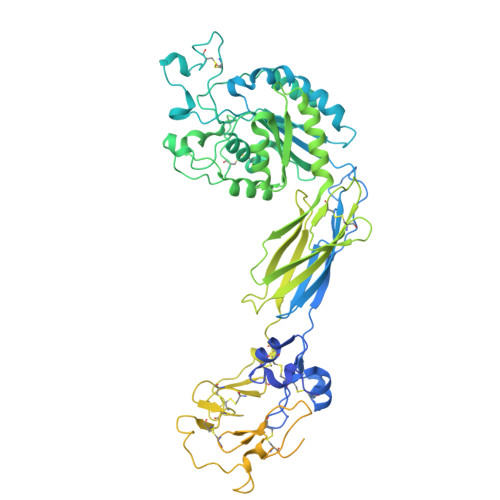
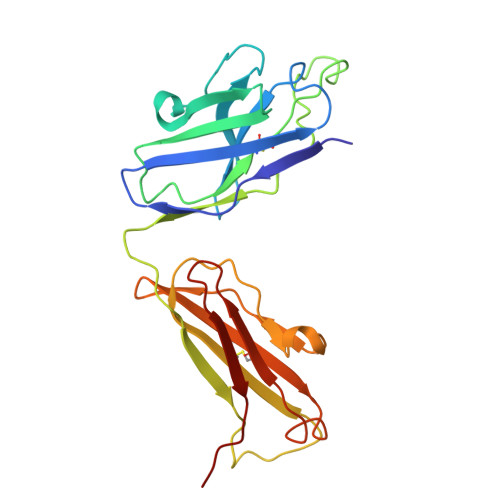
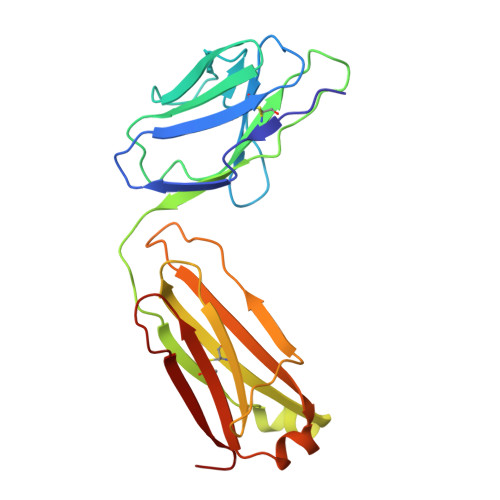
An alpha IIb beta 3 monoclonal antibody traps a semiextended conformation and allosterically inhibits large ligand binding.
Wang, L., Wang, J., Li, J., Walz, T., Coller, B.S.(2024) Blood Adv 8: 4398-4409
- PubMed: 38968144 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1182/bloodadvances.2024013177
- Primary Citation Related Structures:
9AXL - PubMed Abstract:
Monoclonal antibodies (mAbs) have provided valuable information regarding the structure and function of platelet αIIbβ3. Protein disulfide isomerase (PDI) has been implicated in αIIbβ3 activation and binds to thrombin-activated αIIbβ3. Using human platelets as the immunogen, we identified a new mAb (R21D10) that inhibits the binding of PDI to platelets activated with thrombin receptor-activating peptide (T6). R21D10 also partially inhibited T6-induced fibrinogen and PAC-1 binding to platelets, as well as T6- and adenosine 5'-diphosphate-induced platelet aggregation. Mutual competition experiments showed that R21D10 does not inhibit the binding of mAbs 10E5 (anti-αIIb cap domain) or 7E3 (anti-β3 β-I domain), and immunoblot studies indicated that R21D10 binds to β3. The dissociation of αIIbβ3 by EDTA had a minimal effect on R21D10 binding. Cryogenic electron microscopy of the αIIbβ3-R21D10 Fab complex revealed that R21D10 binds to the β3 integrin-epidermal growth factor 1 (I-EGF1) domain and traps an intermediate conformation of αIIbβ3 with semiextended leg domains. The binding of R21D10 produces a major structural change in the β3 I-EGF2 domain associated with a new interaction between the β3 I-EGF2 and αIIb thigh domains, which may prevent the swing-out motion of the β3 hybrid domain required for high-affinity ligand binding and protect αIIbβ3 from EDTA-induced dissociation. R21D10 partially reversed the ligand binding priming effect of eptifibatide, suggesting that it could convert the swung-out conformation into a semiextended conformation. We concluded that R21D10 inhibits ligand binding to αIIbβ3 via a unique allosteric mechanism, which may or may not be related to its inhibition of PDI binding.
- Laboratory of Blood and Vascular Biology, The Rockefeller University, New York, NY.
Organizational Affiliation: