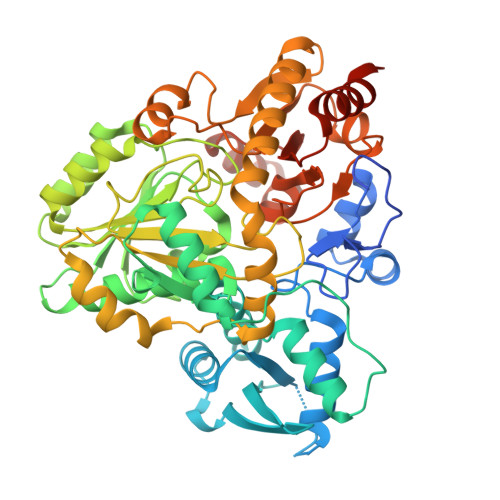
Terminal alkyne formation by a pyridoxal phosphate-dependent enzyme.
Hedges, J.B., Marchand, J.A., Calvo-Tusell, C., Wei, Z.W., Millar, D.C., Garcia-Borras, M., Chang, M.C.Y., Ryan, K.S.(2026) Nat Chem Biol 22: 77-86
- PubMed: 40562852 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-025-01954-9
- Primary Citation Related Structures:
9AUA, 9AUB - PubMed Abstract:
Terminal alkyne-containing natural products can undergo the bio-orthogonal 'click' reaction of Cu(I)-catalyzed azide-alkyne cycloaddition. Recently, an enzymatic mechanism for terminal alkyne formation was discovered in the biosynthesis of L-β-ethynylserine where the pyridoxal phosphate-dependent enzyme BesB forms a rare terminal alkyne-containing amino acid, L-propargylglycine, from a vinyl halide precursor, 4-chloro-L-allylglycine. Here we present the 1.3-Å-resolution crystal structure of BesB with detailed mechanistic and computational studies. We demonstrate that BesB can reversibly catalyze the exchange of the halogen in various 4-halo-allyl-L-glycines, implying the existence of an allene intermediate, which we then also observe. Taken together, this work supports a mechanism whereby an allene is formed from deprotonation-driven halogen loss and the terminal alkyne is then formed by isomerization of the allene. Our work further expands our understanding of the catalytic repertoire of pyridoxal phosphate-dependent enzymes and will enable development of metal-free allene-forming and alkyne-forming biocatalysts.
- Department of Chemistry, University of British Columbia, Vancouver, British Columbia, Canada.
Organizational Affiliation: