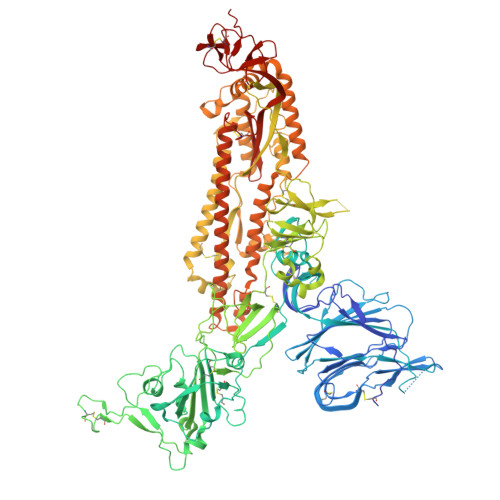
SARS-related coronavirus S-protein structures reveal synergistic RBM interactions underpinning high-affinity human ACE2 binding.
Wang, J., Ma, Y., Li, Z., Yuan, H., Liu, B., Li, Z., Su, M., Habib, G., Liu, Y., Fu, L., Wang, P., Li, M., He, J., Chen, J., Zhou, P., Shi, Z., Chen, X., Xiong, X.(2025) Sci Adv 11: eadr8772-eadr8772
- PubMed: 40085715 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.adr8772
- Primary Citation Related Structures:
8ZY0, 8ZY1, 8ZY2, 8ZY3, 8ZY4, 8ZY5, 8ZY6, 8ZY7, 8ZY9, 8ZYA - PubMed Abstract:
High-affinity and specific binding toward the human angiotensin-converting enzyme 2 (hACE2) receptor by severe acute respiratory syndrome coronavirus (SARS)-related coronaviruses (SARSr-CoVs) remains incompletely understood. We report cryo-electron microscopy structures of eight different S-proteins from SARSr-CoVs found across Asia, Europe, and Africa. These S-proteins all adopt tightly packed, locked, prefusion conformations. These structures enable the classification of SARSr-CoV S-proteins into three types, based on their receptor-binding motif (RBM) structures and ACE2 binding characteristics. Type-2 S-proteins often preferentially bind bat ACE2 (bACE2) over hACE2. We report a structure of a type-2 BtKY72-RBD in complex with bACE2 to understand ACE2 specificity. Structure-guided mutagenesis of BtKY72-RBD reveals that multiple synergistic mutations in four different regions of RBM are required to achieve high-affinity hACE2 binding. Similar RBM changes can also confer hACE2 binding to another type-2 BM48-31 S-protein, which is primarily non-ACE2 binding. These results provide an understanding of how high-affinity hACE2 binding may be acquired by SARSr-CoV S-proteins.
- Guangdong Provincial Key Laboratory of Stem Cell and Regenerative Medicine, Guangdong-Hong Kong Joint Research Laboratory for Stem Cell and Regenerative Medicine, Guangzhou Institutes of Biomedicine and Health, Chinese Academy of Sciences, Guangzhou, China.
Organizational Affiliation: