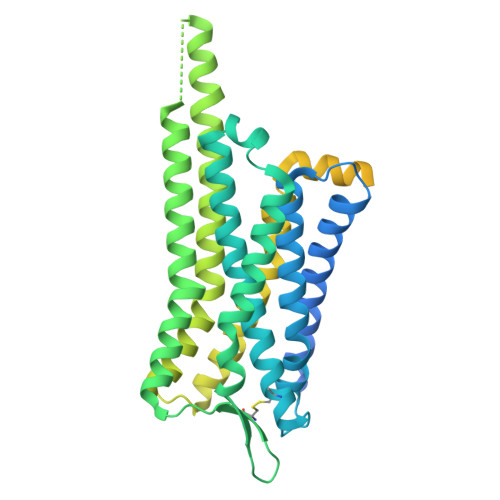
Structural basis for hormone recognition and distinctive Gq protein coupling by the kisspeptin receptor.
Shen, S., Wang, D., Liu, H., He, X., Cao, Y., Chen, J., Li, S., Cheng, X., Xu, H.E., Duan, J.(2024) Cell Rep 43: 114389-114389
- PubMed: 38935498 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2024.114389
- Primary Citation Related Structures:
8ZJD, 8ZJE - PubMed Abstract:
Kisspeptin signaling through its G protein-coupled receptor, KISS1R, plays an indispensable role in regulating reproduction via the hypothalamic-pituitary-gonadal axis. Dysregulation of this pathway underlies severe disorders like infertility and precocious puberty. Here, we present cryo-EM structures of KISS1R bound to the endogenous agonist kisspeptin-10 and a synthetic analog TAK-448. These structures reveal pivotal interactions between peptide ligands and KISS1R extracellular loops for receptor activation. Both peptides exhibit a conserved binding mode, unveiling their common activation mechanism. Intriguingly, KISS1R displays a distinct 40° angular deviation in its intracellular TM6 region compared to other G q -coupled receptors, enabling distinct interactions with G q . This study reveals the molecular intricacies governing ligand binding and activation of KISS1R, while highlighting its exceptional ability to couple with G q . Our findings pave the way for structure-guided design of therapeutics targeting this physiologically indispensable receptor.
- State Key Laboratory of Drug Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, Shanghai 201203, China; University of Chinese Academy of Sciences, Beijing 100049, China.
Organizational Affiliation: