Host genetic variation governs PCV2 susceptibility through CXCL13 and ELK1-mediated immune regulation.
Liu, G., Gao, Y., Cheng, Y., Wang, W., Li, X., Wu, Y., Gao, F., Zhou, Z.W., Sun, Y., Jiang, Y., Yang, N., Shu, Y., Sun, L.(2025) Int J Biol Macromol 310: 143170-143170
- PubMed: 40267997 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2025.143170
- Primary Citation Related Structures:
8ZH9 - PubMed Abstract:
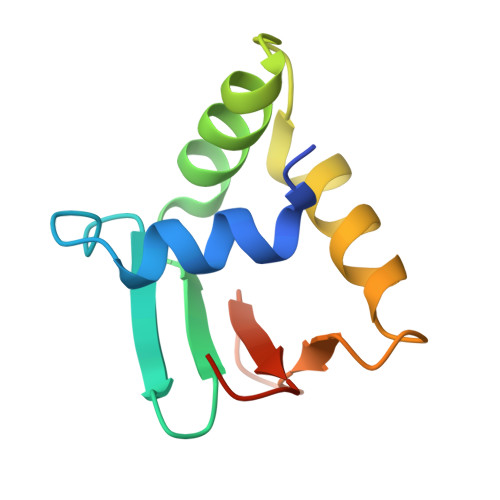
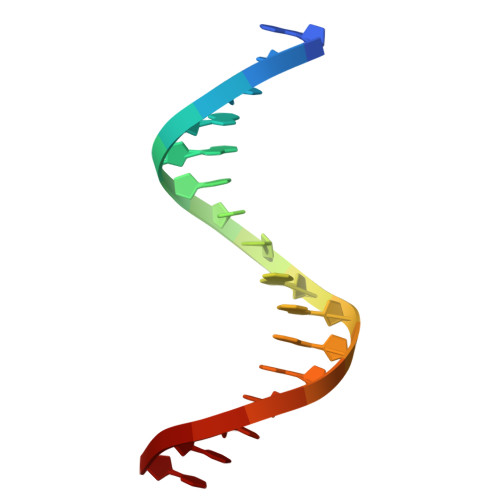
Pathogenic viruses can drive evolutionary adaptations in host biology, leading to diversified immune responses and variable susceptibility among individuals. This study examined how genetic variation in host regulatory regions impacts susceptibility to viral infections. Utilizing a porcine model, we identified the single nucleotide polymorphism (SNP) g.-1014G>A as a critical determinant of CXCL13 expression levels following PCV2 viral exposure. Structural analyses showed that the transcription factor ELK1 specifically recognized and bound to the g.-1014G allele, but not to the g.-1014A allele, through essential residues such as Arg65. This allele-specific binding led to differential CXCL13 expression, with the G allele associated with increased resistance to viral infection. Functional studies demonstrated that CXCL13 played a multifaceted role in antiviral immunity, including the inhibition of viral replication, modulation of immune-related pathways, and attenuation of virus-induced apoptosis. The CXCL13-mediated response involved the activation of the PI3K/Akt pathway, enhancing cell survival during viral challenges. This SNP-dependent regulation of a host factor represented a novel mechanism underlying genetic differences in viral susceptibility, with potential implications for developing broadly applicable antiviral strategies.
- Shenzhen Key Laboratory of Pathogenic Microbes and Biosafety, School of Public Health (Shenzhen), Shenzhen Campus of Sun Yat-sen University, Shenzhen 518107, China; Department of Pathogen Microbiology, School of Basic Medical Sciences, Capital Medical University, Beijing 100069, China.
Organizational Affiliation: